4
Computer and Information Technology in Biomedical and Neuroscience Research
A complex of computer-based resources that can greatly enhance neuroscience research is an attainable goal. Current trends in information technology offer an unprecedented opportunity for neuroscientists to expand their use of hard-won data and to communicate these data more effectively to other scientists. In addition, the sheer mass of neuroscience information accumulated to date and the accelerating rate at which new results are being obtained and reported are becoming major driving forces for the kind of organization, structure, and accessibility that computer-based resources can provide. The attractiveness of the present opportunity is also strengthened by the increasingly intimate role of various computer-based instruments and applications in neuroscience research. This chapter describes how a complex of electronic and digital resources for neuroscience might work in the future, and supports this description with examples obtained in part from the task forces organized to provide advice to the committee and the open hearings the committee sponsored. The chapter also dis-cusses the increasing reliance of biomedical research on computerized resources and the current trends in computer science and information technology that make the goal of a complex of resources attainable.
A neuroscience laboratory of the future will use a variety of computer-based tools
The neuroscience laboratory of the future may seem incredible from our present viewpoint. But a neuroscientist who had fallen asleep over
the microscope in 1970 would awaken today to an equally incredible vision of color graphics workstations, shiny compact disks containing masses of data, and electronic mail messages from colleagues worldwide. Any projections of the future we might generate today are likely to fall short, in some respects, of what will be possible by 2010.
The following scenario illustrates how an experiment might be done in a neuroscience laboratory of the future. (The physiological, anatomical, and biochemical processes it describes are consistent with known aspects of neural function.) Although it is hypothetical and overly simplified, it makes explicit the vision of a family of interrelated databases and other computer-based resources for neuroscience research. Further, it illustrates how three-dimensional graphics, databases, networks, and electronic journals will contribute to research that proceeds more rapidly and efficiently than anything we can imagine today, transcending present conventional information boundaries.
The futuristic scene opens in the laboratory of Jane Smith, who is studying a brain region recently implicated in a particular behavioral disorder that provides a model for human schizophrenia. She uses a new, highly sensitive tract-tracing technique to map this area in rodents that have been genetically engineered to express an altered form of a specific gene. These animals are interesting because they are also deficient in a specific learning task, and this behavioral deficit is quite similar to a behavioral deficit observed in schizophrenic patients. To examine tissue blocks from her experiments for the presence of tracer substances, Dr. Smith uses a microscope connected to a computer workstation, which allows her to record the data onto maps of the brain region and to store all of the data in a digital format. This recording method permits rapid computer visualization of the data in three dimensions.
In her examination, Dr. Smith finds a tiny labeled cell group (area X) embedded in the larger nucleus whose existence has not, to her knowledge, been reported before. While she has the map from her experiment on the video screen, Dr. Smith calls up the three-dimensional anatomical database for this brain region and splits the screen to allow sufficient comparisons of her new map with other maps of the nucleus that have been deposited in the database. She looks at maps from normal animals and maps from genetically engineered animals to see if area X exhibits the same general structure in both groups. One of the maps seems to show a group of cells quite similar to area X. Dr. Smith asks the computer to enlarge that map and show its precise location, compared to her map. (The computer can do this because neuroscientists of the future routinely include with their data
maps certain landmark designations that are used to align and bring into conformity maps from different investigators.)
Dr. Smith finds that the two groups of cells are in precisely the same location and that the structures of the cells match. She then types a message, which is automatically forwarded to the scientist who contributed the map, requesting more information about his work. Later that afternoon, Dr. Smith receives a message from Robert Green, who says he had noticed this cell group in certain kinds of labeling experiments but that it was not his primary interest. Nevertheless, he has attached all the data he has regarding this experiment to his message, with an offer to help in any way he can. Dr. Smith electronically thanks Dr. Green and asks him if he is aware of any chemical characterization of these cells. As the reply is negative, Dr. Smith enters the neurochemical database for this region and pulls up maps showing the distribution of a variety of neurotransmitters. She finds a map of the distribution of a particular protein (zeta) that clearly includes area X and, again, asks for more information. A documentary summary appears at the side of the screen describing the identification of protein zeta by antibody techniques. The summary also states that protein zeta has a specific distribution in the brain and that researchers suspect that it affects potassium channels in the cells' membranes, making the channels less likely to open during synaptic transmission. Dr. Smith transfers the references for these observations into her personal reference files and asks if protein zeta has been sequenced. The reply is that it has been partially sequenced by a Dr. Ungaro in Italy and that the partial sequence is available in a special part of the Protein Identification Resource (PIR) reserved for preliminary data. (Dr. Ungaro, like Dr. Green, is pursuing another interest and has deposited the partial sequence for use by other investigators.)
Dr. Smith enters the PIR and calls up the partial sequence for protein zeta on one side of the screen. On the other side, she retrieves information about the gene that has been altered in her experimental animals. The computer displays the base pair sequence for the gene, followed by the RNA sequence for which the gene codes. From the RNA sequence, the computer displays the expected protein sequence. Dr. Smith instructs the computer to run a comparison of the expected protein sequence with the partial sequence of Dr. Ungaro and discovers a strong match. She immediately types a message to Dr. Ungaro regarding this match and says that she would like to follow up on this information. Dr. Ungaro replies that he is happy that the information was useful and that he will amend the PIR entry to reflect the probable match and her plan to pursue further study.
A few weeks pass, during which Dr. Smith completes another group
of experiments. Her studies show that cells in area X are responsible for the learning deficits in her genetically engineered animals and that protein zeta is an abnormal form that results from the genetic alteration. The protein disrupts the potassium channels by interfering with cyclic AMP, but a new drug that increases the effectiveness of cyclic AMP blocks this effect in Dr. Smith's animals. Moreover, this drug completely alleviates the learning deficits in these animals. Dr. Smith submits these data to an electronic journal, transferring her manuscript text, photomicrographs, sequence comparisons, and other data directly from her computer to the journal. The journal electronically transmits the manuscript to reviewers. Eventually, the manuscript is published in electronic and conventional formats, and the accompanying data are deposited into the appropriate databases. Work in other laboratories then begins to document the biochemical mechanisms that have been discovered and the new drug's effectiveness in human schizophrenics.
As new experiments proceed, Dr. Ungaro's protein is finally sequenced, and this information, along with Dr. Smith's data, is moved by PIR's editors from the preliminary data files of the PIR into its main files of verified data. Dr. Green's contribution is acknowledged in the manuscript and in the anatomical brain database. Such documentation is of great use to investigators who have become interested in protein zeta and its role in schizophrenia. A contribution index database keeps track of the results of data deposits such as those of Drs. Green and Ungaro, and such records are often submitted, along with authored publications, to university tenure and promotion committees. Parts of this scenario could be implemented today; others await further technological, and perhaps sociological, advances. But the useful, coordinated complex of resources suggested in the story will not evolve on its own; it must be planned and put into place. It is the consensus of this committee that now is the time to begin that effort.
Critical Breakthroughs, Important Opportunities
Computers and computer graphics help scientists obtain new images
Neuroscience is an inherently visual science, and in this way it differs from other scientific fields, based on mathematical calculations, that already have benefited from advances in computer science. The widespread use of computers in neuroscience had to wait for major technological breakthroughs in computer graphics, which depended in large measure on increases in the power of individual computers and the associated steep decrease in the cost of that power. In 1945,
it cost approximately $1,000 to perform 1 million computer operations. In 1970, the same number of operations cost less than 6 cents; in 1980, they cost 0.1 cent. By 1997, this cost is expected to decrease by a factor of 100. In terms of time, in 1945, 1 million operations took a month; they took 0.1 second in 1980 and, in 1990, in the case of some of the most powerful workstations, roughly 2.5 milliseconds (U.S. Congress, Office of Technology Assessment, 1990). Such trends are continuing in the present decade.
More than power, the memory capabilities of a computer determine its cost. Again, consistent increases in memory, accompanied by decreases in cost, have occurred over the past three decades (Figure 4-1 ). In 1965, a computer with 64 kilobytes (KB) of memory cost about $200,000 (Bell, 1988) and was used, for example, in expensive satellites and space probes, such as Voyager II. Today, computers of that size cost about $100 and are used to drive popular computer games (e.g., the small, hand-held version of Nintendo®, Gameboy®). Why is a computer capable of running Voyager II necessary for a computer game toy with a tiny video screen? The answer is that the graphics of the toy and the ability to interact with those graphics require far more computer power than is required by calculations (Table 4-1 ). Today's computer graphics—from the dazzling special effects of movies to the variety of the evening news promotions— could not have been accomplished without the recent improvements in computer hardware and software.
Computer graphics is a subspecialty of computer science pioneered by Ivan Sutherland and others in the early 1960s (Goldberg, 1988). The work of these early investigators and the people they trained led to the development of the first primitive graphics workstation less than 20 years ago. The Association for Computing Machinery, one of two major associations of computer science, has a special-interest graphics group that holds a convention every year. In addition to scientific presentations and workshops, this convention includes seminars on various kinds of graphics applications, a computer graphics art show, and exhibits from numerous computer hardware and software companies. The almost 40,000 who now attend this convention, ranging from Hollywood artists to automotive engineers to defense contractors, are an indication of how pervasive, and accessible, computer graphics have become in industry and government (ACM SIGGRAPH, 1989).
The use of computer graphics in biomedical research is also becoming pervasive (albeit not as quickly as in such fields as earth mapping, physics, and space science) and is evolving into the concept of scientific visualization, sometimes called visualization computing (National Academy of Sciences, 1989). In 1987 the National
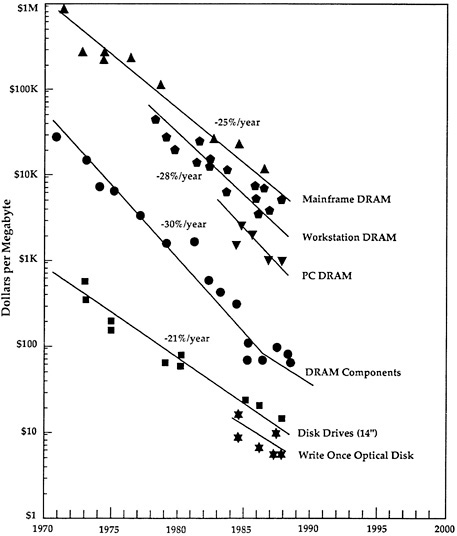
FIGURE 4-1 The cost (in dollars) of computer memory (in megabytes), which has decreased steadily since 1970. DRAM = dynamic random access memory. Chart provided by Mark Duncan of ASKMAR, derived from a graph by Frank Ura of Hewlett-Packard.
Science Foundation convened a panel to consider scientific visualization in many fields, including biomedical science (McCormick et al., 1987). This panel observed that the human eye recognizes geometric and spatial relationships faster than it recognizes other relationships and that the visual display of data would be more efficient for human pattern recognition than displays of numbers and text. Thus, visual-
TABLE 4-1 Text Versus Image Data: Byte Requirements
|
Application |
Example |
Approximate No. of Bytes Required |
|
Word processing |
One computer screen of text, 25 rows × 80 characters / row |
2,000 (2 KB) |
|
Image processing |
||
|
Low-end a |
Magnetic resonance and computed tomography images |
262,144 (256 KB; 100 times more information than a screenful of text) 1,310,720 (1.25 MB) |
|
High-end b |
Computer-aided design (CAD; architecture and engineering applications) |
|
|
Volumetric Imaging c |
Realistic three-dimension- al images with shadows, highlights, reflections, and transparencies |
4,294,967,296 (4 gigabytes; 16,000 times more information than a magnetic resonance or computed tomography image) |
|
Abbreviations: Bit = binary digit, of which there are only two possible: 0 and 1; Byte = the number of bits (usually 8) that represent one character; Kilobyte (KB) = 1,024 bytes; Megabyte (MB) = approximately 1 million bytes; Gigabyte = approximately 1 billion bytes; Terabyte = approximately one trillion bytes. |
||
|
a 512 × 512 pixel display (8 bits/pixel). b1280 × 1024 pixel display (8 bits/pixel). c1K × 1K × 1K voxel display (32 bits/voxel). |
||
ization computing draws on computer graphics, image processing, computer-aided design, signal processing, and user interface studies, with the goal of “transforming the symbolic into geometric reality, [and] offering a method to see the unseen” (McCormick et al., 1987).
One of the most successful applications of scientific visualization in biomedical science has been the modeling of molecular structures from data derived from x-ray crystallography. In the past, such structures were inferred from the numerical coordinate data obtained in complex experiments and then drawn by hand or modeled with sticks and balls to represent atoms or other molecular components. These rep-
resentations depended on the imagination of viewers and their ability to visualize three dimensions from a flat plane and a dynamic image from a static one. Now, computer graphics use mathematical data to generate a moving, dynamic, three-dimensional structure that can change its shape as a consequence of “experimental” conditions entered into the computer or that can be rotated to achieve a different perspective (Plate 4-1 ). These dynamic computer models are sufficiently similar to the in vivo molecules that they may soon help scientists to predict which drugs might stop viruses, how genes are turned on and off, or how two molecules interact with each other (Howard Hughes Medical Institute, 1990). Not only are these models useful for data analysis but they greatly enhance the communication of research results by making those results much more accessible to scientists and nonscientists alike (U.S. Congress, Science Policy Study, 1986).
Virtually every kind of data used in neuroscience can be collected, stored, analyzed, and visualized on a computer. The raw data from some techniques are already collected directly into computer-readable form. For example, in CT, PET, and MRI scans, the data are collected in the computer as numbers, which the computer uses to form images on its monitors. The images can be displayed in a monochrome scale of gray, or they can be enhanced by assigning colors to certain numerical ranges. In all present-day, computer-based image processing, the numerical data rather than the images are stored, because the images can be recreated from numbers at any time. Increasingly, electrophysiological data and data from EEG and ERP studies are also collected in computer-ready form. Later, the traces of electrical activity can be graphically reconstructed on a computer for analysis (Plate 4-2 ).
Anatomical data can be computerized in a variety of ways. For example, electron microscopic photographs of brain tissue can be digitized, and regions of these photographs can be analyzed for total surface area or volume using readily available software programs. One group has reconstructed three-dimensional images of the complex innervation patterns of the cells responsible for body equilibrium from a montage of many electron micrographs (Ross et al., 1990). Sections of whole brains have also been digitized and the sections combined in a three-dimensional display so as to allow rotation and slicing of the brain in any plane of section; information regarding neurochemistry or other variables can then be overlaid on specific regions (Plate 4-3 ; Toga, 1989). An effort to digitize brain sections is under way at the Comparative Mammalian Brain Collection (CMBC), housed at the University of Wisconsin and Michigan State University, to increase the usefulness of this important archival collection (W. Welker, J. Johnson, and S. Greenberg, CMBC, personal communi-
cation, 1990). Beyond its uses in basic science, digitization is also important to clinical neuroradiology. When x-rays are transformed into digital files, the images can be reconstructed in distant locations. This relatively easy method of transmitting important medical records has spurred great interest in building a network system to transmit such images among hospitals (Banks et al., 1986; Elliot et al., 1990).
Until recently, researchers were forced to trace individually stained neurons by hand using a special accessory on a light microscope. Now, however, these drawings can be done automatically with a computerized confocal microscope that reconstructs neurons in three dimensions (Figure 4-2 ). The neuronal images can be studied on their own or used as templates to display differential inputs to specific parts of a neuron. Computer graphics can also be used to display data from pharmacological and neurochemical studies, which normally generate numerical data. The computer can transform numerical measurements, such as dose, response magnitude, and time, into three-dimensional contoured histograms. Such wide-ranging capabilities give investigators unprecedented flexibility in the collection, viewing, analysis, and communication of neuroscience research data.
Increased use of computers in data collection and analysis is apparent from the number of private companies that now market computer hardware and software for neuroscience applications. At the Society for Neuroscience Annual Meeting in 1990, 10 companies exhibited workstations useful for neuroscience research, and 43 companies offered software packages that did almost everything from three-dimensional reconstructions of neurons to quantification and graphic display of fluorescent color changes in ion-sensitive dye experiments. The companies marketing these tools included microscope companies, such as Zeiss and Leitz, as well as traditional computer firms, such as Apple and IBM.
Further evidence of the increasingly important role of scientific visualization in neuroscience comes in the priority given to biomedical computing in universities and government laboratories. For example, Stanford University, Washington University, the University of Pittsburgh, Carnegie Mellon University, and the University of North Carolina, among others, have electrical engineering or computer science departments with sections devoted to the development of technologies, especially graphics and imaging, for biomedical data collection and analysis. Researchers from these departments work in conjunction or in collaboration with counterparts from biomedical departments; some departments also have formal ties to the federally supported supercomputing efforts (Twedt, 1990). At the National Institutes of Health, many individual institutes maintain branches in which com
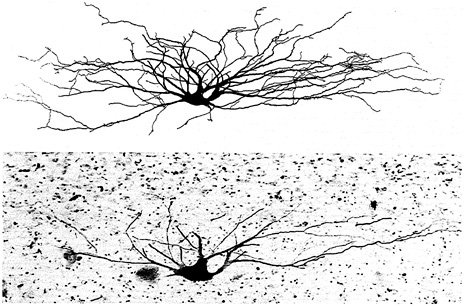
FIGURE 4-2 A spinocervical tract neuron from an adult cat's spinal cord. The lower portion of the figure shows the horseradish peroxidase (HRP)-filled cell that occupies one tissue section. The upper portion is a computer-generated reconstruction from 12 60-micron-thick sections. The cell was identified and stained by M. J. Sedivec, Appalachian State University, and L. M. Mendell, State University of New York, Stony Brook; the reconstruction was performed by J. J. Capowski, Eutectic Electronics, Inc. This photo appeared on the cover of the Journal of Neuroscience, March 1986, and was reproduced with permission from Oxford University Press.
puter scientists and software developers work on applications pertinent to the needs of intramural researchers. For example, when a group of researchers at the National Institute of Mental Health (NIMH) developed a new experimental technique and computer software for visualization and image analysis of brain glucose metabolism, the software was freely shared with any investigator interested in using the new technique (Goochee et al., 1980). NIH also maintains a Division of Computer Research and Technology that, in addition to NIH administrative computing oversight, conducts advanced research into such areas as computer modeling of molecular structure, image processing, and scientific workstation development (National Institutes of Health, Division of Computer Research and Technology, 1989). NIH and the National Science Foundation also provide a number of
grants to develop useful computer tools for scientific research purposes (National Center for Research Resources, 1990; National Science Foundation, 1991). Finally, the National Library of Medicine (NLM) has developed many easily accessible biomedical databases. In addition, the NLM's Center for Biotechnology Information, in conjunction with its efforts to develop useful scientific databases and database linkages, is concerned with the incorporation of computer graphics as user interface tools (National Library of Medicine, 1986).
Considering the primitive state of computer science, especially computer graphics, only 20 years ago, the contributions of these technologies to the visualization of biological processes are quite remarkable. Future work promises to permit scientists to view data in entirely new ways and to integrate those data to a degree never before possible.
Database technologies help organize our knowledge
A major impact of computer technology has been to enable virtually anyone to collect, store, and access many types of data in digital form and to organize data files into discrete units or collections called databases. Although the term database is relatively recent, the concept is as old as symbols carved into clay. The ancient Egyptian records of a grain harvest were databases, as was the census taken by Julius Caesar; the Yellow Pages is a modern example. Computer databases thus reflect the same kind of diversity that other collections of information have shown throughout history. One can classify databases according to what kind of data they contain, how broadly accessible they are, and how formally or informally they are structured (Box 4-1 ).
Databases can be word, number, image, or sound oriented (Williams, 1990). Word-oriented databases are the most prevalent, and the recent growth in their use has been phenomenal. In 1976 there were 750,000 on-line searches of word-oriented databases (e.g., full-text databases, bibliographic databases, and databases containing information synthesized from numerous sources.) In 1988, however, this number rose to 28.3 billion separate queries (Figure 4-3 ). Although less common, the use of number-, image-, and sound-oriented databases is also growing. In 1989 there were more than 5,000 databases of all kinds, which were offered by nearly 900 vendors; they contained nearly 5 billion records (compared with 52 million records in 1975). These statistics do not include the hundreds of thousands of private databases in use by individual research scientists for their own purposes.
In terms of accessibility, databases range from entirely public, such as those available on line in libraries, to entirely private, such as
|
BOX 4-1 RELATIONAL DATABASES VERSUS OBJECT-ORIENTED DATABASES The database management system most widely in use today is the relational system. In a relational database, information is organized in tables. Many of the tables typically have common fields, allowing the user to associate separate tables and to ask questions about the relationships among different pieces of data. A relational neuroscience database, for example, might include the following two tables: a list of drugs, including their behavioral effects and the types of neurotransmitters through which they exert these effects, and a list of anatomical structures, including the predominant cell type in these structures and the neurotransmitter type associated with these structures. Because these two tables have a field in common (the neurotransmitter field), the database user can ask the computer to briefly link the two tables and to generate a list of drugs and their predominant anatomical sites of action. One of the major advantages of a relational system is that each piece of data must only be entered once, thus reducing the possibility of error on the initial entry and when updating is required. Should new research indicate that cocaine, for example, exerts its effects by acting on two neurotransmitter systems instead of only one, this new data can be added to the database and will remain linked to all other pieces of information to which it has a relationship. Relational databases are typically used for textual and not for image data. Database management systems that use object-oriented techniques represent the cutting edge in database technology and are expected to progress rapidly in the coming decade. These systems are designed to support applications with data structures (such as images) that do not fit easily into the table-based structure of the relational system. In an object-oriented system, data need not conform to a standard representation, such as a table, because each piece of data is treated as a separate entity and is stored with its own set of instructions for interacting with other data. This system allows for much greater flexibility in the kinds of data that can be included in a database, and also in the kinds of queries the database can support. |
those kept by an individual scientist for his or her own reference. For example, databases printed out on paper or available on floppy or optical disks (e.g., compact-disk, read-only-memory, or CD-ROMs) can be sold or distributed to the public and thus used in any location. Current Contents, a popular weekly publication listing the tables of contents of selected scientific journals, is now available in computer-readable form on disks. Other databases might be semiprivate or
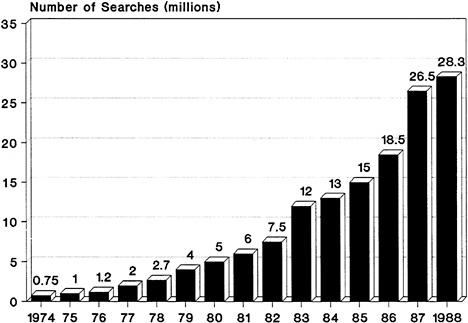
FIGURE 4-3 Growth of on-line searches of word-oriented databases between 1974 and 1988. Adapted from Williams, 1990.
semipublic—shared by a specific group of people—such as several scientists working together on a particular project.
The range of database structures is also broad. Formally structured databases are administered, maintained, updated, and edited in a central location. These databases are generally read-only and include bibliographic reference databases, repositories of historical or archival material, certain types of scientific databases, and directories of various sorts. At the opposite end of the spectrum are informal databases, which are administered, maintained, and updated by users. The user group can be large and membership can be open to everyone, as with the computer bulletin boards that have become so popular. But an informal database can also be structured by a small group of scientists—for example, to keep a “lab notebook” together. Sometimes databases combine both informal and formal components in a unified complex of files. Such a complex might include a bulletin board component, to which users could add data annotations, and a read-only component containing bibliographic reference material. Advances in computer technology make such multicomponent databases feasible by enabling the user to move freely from one com-
ponent to another. Even entirely separate databases are now collectively accessible by software programs that guide users from one database to another. The flexibility to be gained with the different kinds of databases ensures continuing increases in use patterns.
The growth of biomedical databases mirrors the general trends described earlier. Retrieval of information from the scientific literature is basic to the conduct of biomedical research. Yet over the past 30 years, the number of biomedical investigations, especially in the field of neuroscience, has increased substantially, with many of these efforts branching into related subdisciplines. Computer technology was applied to the management of this expansion by the National Library of Medicine, the lead agency for managing and indexing biomedical and scientific literature. In 1964 the NLM created the Medical Literature Analysis and Retrieval System (MEDLARS), a computerized system that catalogs and indexes the library's holdings. In the early 1970s the NLM introduced MEDLINE, which provides on-line access to NLM's records over telephone lines. Using a carefully formulated and controlled list of key words, known as the Medical Subject Headings (MeSH), users can obtain bibliographic reference lists from MEDLINE for a broad range of scientific topic areas.
Today, in addition to MEDLINE, the NLM offers a number of different kinds of on-line databases of scientific information, including information on toxic substances (TOXLINE and its related databases, the Toxicology Data Bank [TDB] and the Registry of Toxic Effects of Chemical Substances [RTECS]) and medical information related to cancer (the Physician Data Query, or PDQ), among others. Databases such as TDB and PDQ, which are often called factual databases, are more than simply bibliographic records because they contain synthesized information from a variety of sources (National Library of Medicine, 1986). The NLM collaborates with other parts of NIH, with universities, and with other research initiatives to develop a broad range of databases for scientific research and medical practice, to establish directories of these resources, and to design useful interfaces among them. In its long-range plan, articulated in 1986, the NLM defined a number of trends for the future, including databases containing image data, full-text storage and retrieval, and electronic publishing of scientific literature.
Especially prominent among the biomedical databases developed outside of the NLM are the protein sequence and genome databases, which serve at least 23,000 scientists (Table 4-2). These databases provide a mechanism by which extensive, complicated sequence and structural data can be compared to what is already known. Comparisons using such databases are many times more efficient than compari-
TABLE 4-2 Selected Genome and Scientific Databases
|
Database |
Date Established |
Kinds of Data |
Size |
|
Protein Identification Resource (PIR) National Biomedical Research Foundation Georgetown University Washington, D.C. |
Early 1970s |
All completely sequenced proteins; aminoterminal sequences and fragmentary sequence data for numerous types of proteins for which sequences are unavailable; bibliographic citations |
9,000 Entries |
|
Protein Data Bank (PDB) Brookhaven National Laboratory Upton, New York |
1973 |
Atomic coordinates and partial bond connectivities as derived from crystallographic studies; structure factor and phase data for some structures; associated textual information and bibliographic citations |
400 sets of atomic coordinates and 100+ sets of structure factors and phases |
|
Genetic Sequence Data Bank (GenBank) Intelligenetics, Inc. Mountain View, California/ Los Alamos National Laboratory Los Alamos, New Mexico |
1979 |
All reported nucleic acid sequences, catalogued and annotated for sites of biological significance in man |
10,000 Records |
|
On-line Mendelian Inheritance in Man (OMIM) Welch Medical Library Johns Hopkins University Baltimore, Maryland |
1987 |
Full-text and bibliographic information on known inherited human traits and disorders; linked to gene map and molecular defects in Mendelian disorders list; linked to GDB |
Data on more than 5,000 traits and disorders |
|
Chemical Abstracts Service Source Index (CASSI) American Chemical Society Chemical Abstracts Service (CAS) Columbus, Ohio |
Early1970s |
Complete bibliographic descriptions of more than 60,000 serials and nonserials monitored by the CAS (1830–present); guides to depositories of unpublished data; holdings information for numerous U.S. libraries and several libraries abroad |
400 Billion bytes |
|
Genome Data Base (GDB) Welch Medical Library Johns Hopkins University Baltimore, Maryland |
1990 |
Mapping database–contains chromosome band locations, genes, anonymous DNA segments, fragile sites, DNA probes, and links to source data (e.g., bibliographic references, personal communications, and conference abstracts); extensions to include genetic linkage and order information from various mapping methods; linked to OMIM |
Data on 2,000 genes, 6,000 mapped D-segments, and 20,000 references |
|
SOURCE: Vela, 1990, and Williams, 1990. |
|||
sons done by hand and have resulted in new discoveries by database users (see Box 2-1 ). These databases are also a valuable source of experience and knowledge for those involved in developing analogous resources for neuroscience. For example, each of the protein sequence and genome databases was developed independently, often by an individual scientist, and each has a different history of funding, administration, oversight, software development, and data organization (Smith, 1990; Vela, 1990). Although efforts are now under way to integrate these databases, such integration is not without considerable difficulty and expense.
Ideally, users should be able to access information from a number of protein or genome databases, but currently this is impossible because independent development has resulted in quite separate database structures. For example, some of the early databases (e.g., the Protein Identification Resource [PIR] and the Protein Data Bank [PDB]) were developed from isolated files of information without the use of a database management system for indexing and retrieving entries. This strategy worked only as long as the number of entries was relatively small and the databases stood alone. Present efforts to link the PIR and PDB must first establish a database management system for each and create links between the two database environments. The National Center for Biotechnology Information at the NLM is devising linkage software that will make PIR and the Genetic Sequence Data Bank (GenBank) accessible by similar means and commands. GenBank has also been working to strengthen its links to the European Molecular Biology Library (EMBL) and the Japanese DNA Data Bank. In contrast, the Genome Data Base (GDB) at Johns Hopkins has planned for linkages to other databases from its inception.
The databases also display important differences in software development and the availability of analytical tools. PIR includes some analytical software, whereas PDB makes only the code available for analysis, which often leaves the user confronted with unfamiliar software. Other databases, such as GenBank, spend up to 50 percent of their budgets to develop software for analytical purposes and for database conversion to more modern management systems. Making these conversions is time consuming and costly, but also necessary to extract the most useful information from these resources.
The protein sequence and genome databases reflect a variety of funding mechanisms. GenBank, a collaborative project of Los Alamos National Laboratory and Intelligenetics, Inc., is supported by the broadest range of sources, including the National Science Foundation (NSF); the Departments of Defense, Energy, and Agriculture; and NIH, through the NLM, the National Center for Research Resources,
and the National Institute of General Medical Sciences. PDB receives funds from NSF, NLM, and DOE. The Howard Hughes Medical Institute, a private agency, currently funds the GDB and the On-line Mendelian Inheritance in Man (OMIM) database. However, in the near future, GDB and OMIM will be supported by federal funding.
In an effort sponsored in part by the NSF and NLM, the Corporation for National Research Initiatives is developing an experimental National Digital Library System (DLS). The initial target databases are MEDLARS and the GDB and OMIM systems at Johns Hopkins University. DLS will network these databases and employ “knowledge robots” (knowbots®)1 to initiate and manage complex, multiple-database searches. This system makes it unnecessary for users to know any details of how to search a database, where the database is located, or even how many different databases exist (Cerf and Kahn, 1988).
It is clear that much can be learned from the experiences of the protein and genome databases. But as noted in the previous chapter, neuroscience data differ significantly from protein and gene sequence data and present complex problems in terms of three-dimensional displays and integration across varied data types. Despite the growing use of database technology in individual neuroscience laboratories, relatively few steps have been taken to construct databases for general use by neuroscientists. Understandably, most neuroscience databases either target specific regions of the brain, or are designed as an organizational system (to which data can be added by the user) or as a method for making archival data more accessible. Some of the neuroscience database development efforts receive targeted funding; others do not. These efforts are important because they demonstrate a growing scientific need and address the various kinds of data inherent in neuroscience, especially images. We describe two such efforts below.
A neuroscience information management system called Brain Browser has been developed for use on Macintosh computers by a team of neuroscientists with computer expertise (Bloom, 1989). The system is based on HyperCard ®, which organizes information in a relational manner, similar to cross-referenced index cards. Contained within the system is information regarding 300 brain regions in the rat, including 900 separate circuits, and references. In addition, the system includes a complete stereotaxic atlas for the rat brain, which can be used as a graphic-based interface to access circuitry data (Figure 4-4 ). Rather than being a self-contained, read-only system, Brain Browser can be expanded by users, who may add “index cards” of their own references and data and customize the database for their own use and that of others (if they share their entries).

FIGURE 4-4 A typical computer screen from the Brain Browser software program. Figure courtesy of Floyd Bloom, Scripps Clinic and Research Foundation. Reprinted with permission of Raven Press.
Another group of neuroscientists and computer scientists from three universities is developing computer tools for high-resolution capture, storage, and organization of neuroscience data (Hillman et al., 1990). Their objective is to construct three-dimensional color atlases, based on real images of brain sections, that are linked to other image, text, and numerical data regarding the anatomy, chemistry, and physiology of specific brain regions. This work is also intended to develop useful software for the extraction of important features and for interfaces that allow transitions from one data type to another. All efforts are focused on one brain region, the thalamic nuclei. Such a focus allows the investigators, who are already expert in thalamic function, to assess more adequately the usefulness and feasibility of the tools being developed.
These two examples may obscure the fact that the development of useful tools to manage neuroscience information is really in its embryonic stages. The majority of neuroscientists use computers for word processing or to organize references and, sometimes, their data images. These are highly individual approaches and vary widely in sophistication. The next section on networks argues that electronic transmission of data sets, including images, most likely will be an
integral part of science in the future. Whether neuroscience will be ready for this kind of future depends on the actions taken now to develop the ability to organize data and combine that organization with data collection.
Electronic networks greatly increase communication
The scientific enterprise is composed of men and women who generate ideas, design ways to test those ideas, collect data, and communicate the ideas and data in a variety of ways. The communication of ideas and results is as important to the growth of knowledge as the data themselves. Scientists meet at formal gatherings, discuss experiments with their colleagues, publish papers, and talk to each other by telephone and electronic mail. One of the major goals of computer network development is to create a communication environment that is as free of barriers as possible—an environment that can support the rapid communication of ideas and images at every stage of experimentation and discovery. Some have envisioned this environment as a “National Collaboratory”2 (Lederberg and Uncapher, 1989). Its cornerstone would be vast networks of electronic links built on a foundation that was begun little more than 20 years ago.
The first multipurpose wide area network (WAN) was developed in 1969 to link computers at 30 research campuses whose work was supported by the Department of Defense. Called ARPANET, the network allowed the sharing of large, expensive computers and facilitated electronic message delivery among widely scattered university and industrial research sites throughout the United States. ARPANET was extremely successful because of a specially developed mechanism for routing messages, called packet switching. Packet switching is unlike the circuit-switching mechanism that connects one telephone to another; no actual circuit is established when a message is sent over a packet-switched network. Rather, the messages are broken up into packets, like envelopes, with address information attached. These packets move from one network node to another, as a letter moves from post office to post office until it reaches its destination. Electronic messages arrive within fractions of seconds, however, not days (Kahn, 1987).
Local area networks (LANs) link computers across short distances (within an office building or among university laboratories, for instance) and are often linked themselves to national and international WANs. Since their introduction in the 1970s, more than 5 million LANs have been installed. In 1973, the Defense Advanced Research Projects Agency (DARPA), which funded the development of ARPANET 's
packet-switching technology, began a project to develop technology to interconnect different packet-switching networks. A series of computer communication protocols were developed for this purpose, along with “gateways” to interlink different LAN and WAN computer networks. The so-called TCP/IP protocol suite that resulted is used today to interconnect 5,000 networks in 35 countries supporting more than 300,000 computers that range from supercomputers to workstations and even personal computers. The system, which is called the Internet (Quarterman, 1989), is operated on a collaborative, cooperative basis involving government, military, university, and industrial resources and thousands of volunteers who keep the system operating.
In 1988, the National Science Foundation began work on the NSFNET to interconnect a small number of supercomputer centers and to sponsor the creation of intermediate-level, or regional, networks to provide access to the NSFNET and its supercomputing resources. Some 13 supercomputer centers and 400 universities are linked to the NSFNET, along with more than 2,000 other networks whose traffic the NSFNET supports. The NSFNET and its “entourage” are now an integral, vital component of the international Internet.
The full utility of such large-scale networking can be realized only if standards are established to enable effective communication among the computer systems that constitute the network (National Academy of Sciences, 1989). The Internet Activities Board and its subsidiary groups, the Internet Research and Engineering Task Forces, are responsible for guiding research and development of standard protocols for Internet computer communication. The quest for useful, yet practical, standards for the communication of numeric, text, image, and other signals data continues and could play an important role in the creation of an effective national neural circuitry database.
One of the most critical limitations of networks for scientific applications is the capacity, or bandwidth, of the present links. Bandwidth refers to the amount of data that can be transmitted per second. For example, NSFNET began with links capable of transmitting 56 kilobits of information per second. A full-text article from a scientific journal averages 20 kilobytes, or 160 kilobits; an abstract averages 16 kilobits. Thus, NSFNET's initial bandwidth was sufficient to transmit three to four abstracts, but not even one entire journal article, per second. In 1989, NSFNET upgraded its system to links capable of transmitting 1.5 megabits per second—large enough for 10 full-text articles or approximately 100 abstracts. Upgrades now in progress will increase the NSFNET backbone-link bandwidth to 44.7 megabits per second, almost a 30-fold increase in the amount of information transmitted every second. Traffic in the NSFNET backbone has reached
nearly 5 billion packets per month, and the rate of increase is still substantial. The introduction of 45 megabit-per-second capability into the NSFNET is an intermediate step in the race toward gigabit-(a billion bits)-per-second service, which will be needed as communication of image data becomes more prevalent.
A single line drawing or simple black-and-white picture represents the same amount of data as an entire journal article of text. An uncompressed transmission rate of 180 megabits per second is required to run a color movie on a conventional TV screen in real time. Computer-generated images on high-resolution screens require much more: 1.44 × 109 bits per second. Consequently, for useful multimedia network transmission, transmission rates will have to reach at least the gigabit-per-second range or higher. (A gigabit equals the information contained in 80,000 double-spaced pages of text.)
The importance of building a nationwide high-bandwidth network is underscored by the attention such a network is receiving from policymakers. An early proposal for a nationwide system of electronic links among supercomputers and libraries was made by Senator Albert Gore in 1979, and the resulting legislation was considered by Congress in 1990 and is being reintroduced this year (Gore, 1990). In 1985, Congress passed Senator Gore's Supercomputer Network Study Act, authorizing the Office of Science and Technology Policy (OSTP) to analyze the need for such a network. Citing the lack of adequate network technology for “scientific collaboration or access to unique scientific resources ” and the aggressive efforts of Japan and Europe to upgrade their networks, the OSTP report, issued in 1987, strongly recommended that federal resources be allocated to upgrade existing networks and undertake the research necessary to advance network technology even farther. In 1988, the National Research Council's National Research Network Review Committee issued a report that expanded the specific recommendations made by OSTP but also strongly recommended the implementation of a National Research Network. This recommendation was strongly endorsed in 1989 by the National Academy of Sciences' Panel on Information Technology and the Conduct of Research (National Academy of Sciences, 1989). A further indication of the attention being devoted to this issue was a report entitled The Federal High Performance Computing Program, delivered to Congress in 1989 by the President's science advisor, D. Allan Bromley (Office of Science and Technology Policy, 1989). This program plan again recommended a phased development of a National Research and Education Network, now commonly called NREN. Three major objectives specified by all of these initiatives were (1) to upgrade the bandwidth of the backbone networks currently serving
science and education to at least gigabit-per-second speed beginning in the mid- to late 1990s; (2) to develop appropriate technology and communication standards that would significantly improve the ability to communicate over electronic networks; and (3) to increase the ubiquity of networking (the NREN program, in particular, proposes to establish connectivity between 1,000 research and education institutions).
Some electronic collaboratory groups are forming now
A critical mass of enthusiasm has been building among legislators, policymakers, and scientists about the possibilities offered by network technology to provide an infrastructure for enhanced scientific collaboration. An interesting project to realize the collaboratory concept is now under way (Schatz, 1991). The project's objectives are to encode, in digital form, all knowledge of the community of biologists who study the worm Caenorhabditis elegans and to construct an integrated computer environment to manipulate this knowledge across the Internet. There are about 500 scientists in this multidisciplinary “worm community,” representing genetics, anatomy, physiology, biochemistry, and neuroscience, and the community has always had a high degree of openness in communication and data sharing. For example, for more than a decade, the group has maintained a newsletter of news and research findings to which anyone may contribute. On numerous occasions, the newsletter has reported significant research in advance of formal publication, thus functioning as an electronic bulletin board.
Genetic mapping is a major aim of the worm community, as it is of many scientists engaged in the study of invertebrate models (National Research Council, 1985). Alan Coulson and John Sulston have already established a physical map database of C. elegans genes, using DNA fragments that they made or that were sent to them by other researchers (Coulson et al., 1986). About 70 percent of the worm's 100 million-base-pair genome has been mapped thus far. This information, updated monthly through Bitnet and Internet, is contained within a digital database that is distributed to several sites; from there, the database can be accessed by modem or direct electronic connection by any worm laboratory.
The current initiative intends to draw on the successful experience of the physical map database and newsletter to build an electronic worm community. Plans include making the entire body of C. elegans knowledge available in digital form and constructing strong electronic platforms for informal communication and to support powerful
analysis and annotation, as well as retrieval. This project in many ways could be a “test bed” for neuroscience because it involves the conversion of many types of data into digital forms suitable for inclusion in a coordinated, multimedia research resource. For example, so much is known about the neurons that make up the worm's nervous system that each neuron's developmental lineage, wiring diagram, and physical location are understood. The organization of this neural information into a mixed text-numerical-image database would be of tremendous value to future work with data from more complicated organisms.
The openness of the worm community offers another important model for developers of computerized resources for neuroscience because a major portion of the current project involves the inclusion of informal data in the database and the establishment of electronic means of communication. These informal data include everything from recipes for laboratory reagents to intermediate experimental results. How the various kinds of informal data are handled, how continued openness is encouraged and rewarded, and how interfaces are designed for the most efficient use of resources by the community will be applicable in the larger context of neuroscience. Most important, because the worm community cannot answer all the questions posed by the incorporation of these technologies into the field ofneuroscience, consideration of the worm community's efforts will help to focus the design of specific pilot projects in the larger neuroscience context.
Conclusion
The role of electronic networks in biomedical science can be expected to expand as network capabilities are upgraded. With these upgrades and the implementation of such projects as the National Digital Library System and the worm community initiative, the formation of national or international collaboratories becomes a more realistic goal for many fields of science. For neuroscience, high-resolution computer imaging and high-bandwidth networking are converging, and major obstacles to the integration of enabling technologies in the research enterprise are quickly diminishing. By beginning now, the integration of these technologies into neuroscience will be ensured. As an added benefit, integrating these technologies into a field as complex as neuroscience will help to generate new ways of conducting scientific exploration that will be applicable to all of biomedical and biological science.
References
ACM SIGGRAPH. 1989. Computer Graphics: SIGGRAPH ‘89 Conference Proceedings 23(3).
Banks, G. , J. K. Vries , and S. McLinden. 1986. Radiologic automated diagnosis. IEEE 195 : 228-239.
Bell, C. G. 1988. Toward a history of (personal) workstations. Pp. 1-47 in A History of Personal Workstations , A. Goldberg , ed. New York : ACM Press.
Bloom, F. 1989. Databases of brain information. Chapter 13 in Three-Dimensional Neuroimaging , A. Toga , ed. New York : Raven Press.
Capowski, J. J. , M. J. Sedivec , and L. M. Mendell. 1986. An illustration of spinocervical tract cells and their computer reconstruction (cover). Journal of Neuroscience 6(3).
Cerf, V. , and R. Kahn. 1988. The Digital Library System: The World of Knowbots , vol. 1. Reston, Va. : Corporation for National Research Initiatives. Coulson, A. , J. Sulston , S. Brenner , and J. Karn. 1986. Towards a physical map of the genome of the nematode Caenorhabditis elegans . Proceedings of the National Academy of Sciences USA 83 : 7821-7825.
Elliot, L. P. , S. K. Mun , S. C. Horii , and H. Benson. 1990. Digital Imaging Network System (DINS) Evaluation Report. Washington, D.C. : Department of Radiology, Georgetown University Medical Center.
Goldberg, A. , ed. 1988. A History of Personal Workstations. New York : ACM Press.
Goochee, C. , W. Rasband and L. Sokoloff. 1980. Computerized densitometry and color coding of [14C] deoxyglucose autoradiographs. Annals of Neurology 7(4) : 359-370.
Gore, A. 1990. Networking the future. Washington Post, July 15.
Hillman, D. E. , R. R. Llinas , M. Canaday , and G. Mahoney. 1990. Concepts and methods of image acquisition, frame processing, and image data presentation. Pp. 3-38 in Three-Dimensional Neuroimaging , A. Toga , ed. New York : Raven Press. Howard Hughes Medical Institute. 1990. Finding the Critical Shapes. Bethesda, Md. : Howard Hughes Medical Institute Office of Communications.
Kahn, R. E. 1987. Networks for advanced computing. Scientific American 257 : 136-143.
Lederberg, J. , and K. Uncapher , co-chairs. 1989. Towards a National Collaboratory: Report of an Invitational Workshop. Rockefeller University, New York City , March 17-18.
McCormick, B. H. , T. A. DeFanti , and M. D. Brown , eds. 1987 Visualization in scientific computing. SIGGRAPH Computer Graphics Newsletter 21(6) : 1-13.
National Academy of Sciences. 1989. Information Technology and the Conduct of Research. Washington, D.C. : National Academy Press.
National Center for Research Resources. 1990. Biomedical Research Technology Resources: A Research Directory. Pub. No. 90-1430. Bethesda, Md. : National Institutes of Health.
National Institutes of Health , Division of Computer Research and Technology. 1989 Annual Report. Bethesda, Md. : National Institutes of Health.
National Library of Medicine. 1986. Obtaining Factual Information from Data Bases: Report of Long Range Planning Panel 3. Bethesda, Md. : National Institutes of Health.
National Research Council. 1985. Models for Biomedical Research: A New Perspective. Washington, D.C. : National Academy Press.
National Research Council. 1988. Toward a National Research Network. Washington, D.C. : National Academy Press.
National Science Foundation. 1991. Guide to Programs. Pub. No. 038-000-00585-5. Washington, D.C. : U.S. Government Printing Office.
Office of Science and Technology Policy , Executive Office of the President. 1987. A Research and Development Strategy for High Performance Computing. Washington, D.C. November 20.
Office of Science and Technology Policy , Executive Office of the President. 1989. The Federal High Performance Computing Program. Washington, D.C. September 8.
Quarterman, J. S. 1989. The Matrix. New York : Digital Press.
Ross, M. D. , L. Cutler, G. Meyer, T. Lam , and P. Vaziri. 1990. 3-D components of a biological neural network visualized in computer generated imagery. Acta Otolaryngologia (Stockholm) 109 : 83-92.
Schatz, B. R. 1991. Building an Electronic Scientific Community. Pp. 739-748 in Proceedings of the 24th Annual Hawaii International Conference on System Sciences , IEEE Computer Society , vol. 3.
Smith, T. F. 1990. The history of the genetic sequence databases. Genomics 6 : 701-707.
Toga, A. , ed. 1989. Three-Dimensional Neuroimaging. New York :.
Twedt, S. 1990. Biologists find speed, imaging powers of supercomputers key to research The Scientist (Aug. 20) : 10-11.
U.S. Congress , Office of Technology Assessment. 1990. Critical Connections: Communication for the Future. OTA-CIT-407. Washington, D.C. : U.S. Government Printing Office.
United States Congress, Science Policy Study. 1986. The Impact of the Information Age on Science. Hearings, vol. 10. Pub. No. 57-095O. Washington, D.C. : U.S. Government Printing Office.
Vela, C. 1990. Overview of U.S. Genome and Selected Scientific Databases. Background paper prepared for the Committee on a National Neural Circuitry Database , Institute of Medicine.
Williams, M. E. 1990. The state of databases today: 1990. Pp. vii-xiv in Computer-readable Databases: A Directory and Data Sourcebook , K. Y. Marcaccio and J. Adams , eds. Detroit, Mich. : Gale Research, Inc.
Notes
1. Knowbots is a registered trademark of the Corporation for National Research Initiatives.
2. The concept of a national collaboratory does not ignore the essentially international character of science. Responsibility for the infrastructure of each national collaboratory should rest, as it traditionally has, with the governmental agencies of each nation that places a priority on scientific excellence. The international collaboratory would be created simply by establishing links among these national resources.

























