This paper was presented at a colloquium entitled “Genetic Engineering of Viruses and of Virus Vectors,” organized by Bernard Roizman and Peter Palese (Co-chairs), held June 9–11, 1996, at the National Academy of Sciences in Irvine, CA.
Negative-strand RNA viruses: Genetic engineering and applications
PETER PALESE *, HONGYONG ZHENG, OTHMAR G. ENGELHARDT, STEPHAN PLESCHKA, AND ADOLFO GARCÍA-SASTRE
Department of Microbiology, Mount Sinai School of Medicine, 1 Gustave L.Levy Place, New York, NY 10029
ABSTRACT The negative-strand RNA viruses are a broad group of animal viruses that comprise several important human pathogens, including influenza, measles, mumps, rabies, respiratory syncytial, Ebola, and hantaviruses. The development of new strategies to genetically manipulate the genomes of negative-strand RNA viruses has provided us with new tools to study the structure-function relationships of the viral components and their contributions to the pathogenicity of these viruses. It is also now possible to envision rational approaches—based on genetic engineering techniques—to design live attenuated vaccines against some of these viral agents. In addition, the use of different negative-strand RNA viruses as vectors to efficiently express foreign polypeptides has also become feasible, and these novel vectors have potential applications in disease prevention as well as in gene therapy.
DNA-Containing Viruses
Among animal viruses, DNA-containing viruses were the first to become amenable to genetic engineering techniques. This breakthrough was achieved for simian virus 40 when a cloned cDNA copy was transfected into cells, resulting in the formation of infectious virus (see Table 1). Transfected mutated cDNA molecules gave rise to defined mutant viruses (1). A second methodology involving the use of homologous recombination allowed, for the first time, the rescue of large DNA-containing viruses such as herpes viruses (2). In this approach, intact herpes viral DNA as well as cloned DNA flanked by viral sequences was transfected into cells. Homologous recombination between the cloned DNA and the wild-type genome can occur, and novel viruses can be selected under appropriate conditions. For example, recombinants with DNA fragments containing a viral thymidine kinase gene can be selected in appropriate cell lines and media, and viruses lacking a thymidine kinase can be isolated in the presence of nucleoside analogs (e.g., Ara T). This general technique allows the successful construction of viral variants of herpes viruses, and similar procedures have been developed for pox viruses (3, 4) and other DNA-containing viruses including adenoviruses (5) and parvoviruses (6). Finally, strategies have been developed to generate infectious as well as mutant viruses by transfecting cosmids containing overlapping portions of large viral genomes. Viruses arise via recombination between the cosmids. This system was successfully used to rescue infectious herpes simplex 1 viruses (7), cytomegaloviruses (8) and Epstein-Barr viruses (9) from their respective cosmids.
Positive-Strand RNA Viruses
RNA-containing viruses belong to a variety of families with diverse replication strategies. Unique among the RNA viruses are the retroviruses, whose replication involves a double-stranded DNA phase, making these viruses an easy target for genetic manipulation. Transfection of full-length cDNA molecules leads to the establishment of replicating virus particles and integration of the viral genetic information into the host genome (10). The engineering of retroviral genomes has become one of the most successful genetic approaches in modern virology and is central to the study both of viral gene expression and of protein structure-function analysis. In addition, retrovirus constructs are among the most widely used vectors for gene transfer and gene therapy (11).
Most of the other positive-strand RNA viruses are also amenable to genetic engineering approaches (Table 1). In the case of the small and medium sized positive-strand RNA viruses, full-length genomic RNA has been shown to be infectious when transfected into cells. Plus-strand RNA serves as mRNA for the synthesis of viral proteins as well as template for viral RNA replication. Thus, transfection of cloned DNA of poliovirus RNA (or of cDNA-derived RNA) into permissive cells results in the formation of infectious virus particles (12).
Remarkably successful have been studies using Sindbis viruses and Semliki forest virus (13, 14). The cDNA-derived RNAs of these positive-strand RNA viruses can be used to efficiently rescue infectious viruses, thus allowing an extensive analysis of the promoter elements of the viral RNAs as well as structure-function studies of the viral proteins. Furthermore, these viruses have received increased attention because of their potential for expressing copious amounts of heterologous genes via recombinant constructs. Up to 108 molecules of heterologous protein per cell have been expressed using these systems.†
Introduction of cDNA-Derived RNA into a Negative-Strand RNA Virus (Influenza Virus)
The life cycle of negative-strand RNA viruses differs from that of the other RNA viruses in many ways. Specifically, the genomic RNA of negative-strand RNA viruses is not infectious, and infectious virus particles must also deliver their own RNA-dependent RNA polymerase into the infected cell to start the first round of virus-specific mRNA synthesis.
Thus, approaches different from those used for positive-strand RNA viruses had to be developed to allow the rescue of
The publication costs of this article were defrayed in part by page charge payment. This article must therefore be hereby marked “advertisement” in accordance with 18 U.S.C. §1734 solely to indicate this fact.
Abbreviations: RNP, ribonucleoprotein; HA, hemagglutinin; NA, neuraminidase, VSV, vesicular stomatitis virus.
|
* |
To whom reprint requests should be addressed, e-mail: ppalese@smtplink.mssm.edu. |
|
† |
Belli, B.A., Polo, J.M., Driver, D.A., Latham, E., Banks, T.A., Chang, S.M.W. & Dubensky, T.W., Jr., National Academy of Sciences Colloquium on Genetic Engineering of Viruses and of Virus Vectors, June 9–11, 1996, Irvine, CA, no. 1. (abstr.). |
Table 1. Genetic engineering of animal viruses
genetically engineered viruses of these virus families (Table 1). Site-specifically altered influenza viruses were first obtained by reconstituting in vitro a biologically active ribonucleoprotein complex (made of synthetic RNA and purified nucleoprotein and polymerase proteins) and then transfecting the complex into helper virus-infected cells (Fig. 1) (15). The helper virus provides in trans the viral proteins required for amplification of the synthetic RNP complex. Subsequent reassortment of the synthetic gene and helper virus-derived RNA segments, followed by selection for the reassortant (transfectant) virus, allows the introduction of site-specific changes into the genome of influenza viruses (16). Selection of the transfectant virus can be achieved by choosing host range or temperature-sensitive mutants as helper viruses. Alternatively, antibody preparations specific for the viral surface proteins can be used to select against the helper virus or for these novel viral constructs. Following such protocols, six of the eight genes [PB2, hemagglutinin (HA), neuraminidase (NA), NP, M and NS] of influenza A viruses and the HA of an influenza B virus have now successfully been altered by genetic engineering methods (17–22).
Plasmid-Based Reverse Genetics System for Influenza Virus
A method was recently developed to reconstitute a biologically active influenza virus RNP complex within a cell rather than in vitro. This alternative approach avoids the need to purify viral proteins and to transfect an RNA-protein complex into cells; instead, this method involves the transfection of plasmids. The first plasmid contains a human polymerase I promoter and a hepatitis delta virus-derived ribozyme sequence which flank the synthetic influenza virus gene. The polymerase I-driven plasmid is cotransfected into human cells with polymerase II-responsive plasmids expressing in trans the viral PB1, PB2, PA, and NP proteins. Such a system involving the use of five plasmids allows the amplification and expression of a synthetic influenza virus gene and takes advantage of the convenience of plasmid transfections as compared with RNP transfections (23). Using this approach, it was possible to rescue a synthetic NA gene into a recombinant influenza A virus. A synthetic HA gene has also been rescued by this novel technique (Fig. 2) (A.G.-S., unpublished results). It should be noted, however, that this plasmid-based reverse genetics system still relies on the presence of a helper virus which provides the genetic backbone into which the plasmid-derived gene can be introduced.
Chimeric Influenza Viruses Expressing Foreign Epitopes or Polypeptides
The development of methods to rescue synthetic RNAs into the genomes of influenza viruses allowed the construction of chimeric viruses expressing a variety of foreign epitopes. Specifically, epitopes derived from HIV, plasmodia, or lymphocytic choriomeningitis virus proteins were successfully expressed in either the HA or the NA of different influenza viruses (16, 24). Such constructs were shown to induce a potent B-cell and/or T-cell response against the foreign epitope in experimental animal systems. Specifically, Li et al. (25) gen-
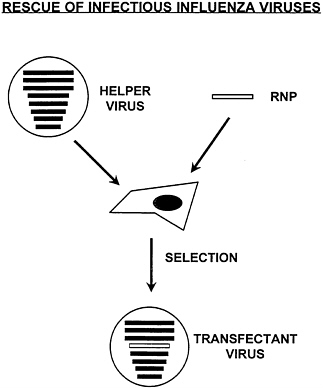
FIG. 1. A reverse genetics system for the rescue of infectious influenza viruses containing cDNA-derived RNA. The method allows the substitution of one of the eight genomic RNA segments of the virus by a synthetic RNA. A biologically active viral ribonucleoprotein complex (RNP) is made in vitro by mixing cDNA-derived RNA with purified viral nucleoprotein and polymerase proteins. The RNPs are transfected into cells which have been previously infected with an influenza helper virus. Using a selection method, viruses containing the genetically engineered RNP (transfectant viruses) can be isolated.
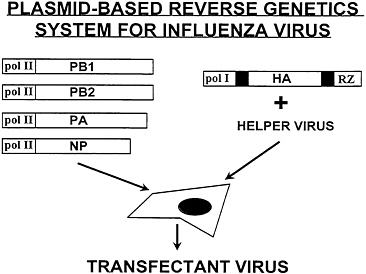
FIG. 2. A plasmid-based reverse genetics system for the rescue of infectious influenza viruses containing a genetically engineered segment. Cells are transfected with four plasmids that are able to express the viral NP and polymerase (PB2, PB1, and PA) proteins from a cellular polymerase II-responsive promoter (pol II). An additional plasmid which contains, for example, the HA open reading frame flanked by the 5′ and 3′ noncoding regions of the viral RNA segment (black boxes) is cotransfected. The HA plasmid is able to express an HA-specific viral RNA by transcription from a polymerase I-responsive promoter (pol I) followed by the ribozyme (RZ)-mediated cleavage of the transcript. The HA-specific RNA segment is intracellularly complexed with the NP and polymerase proteins to form RNPs that can be rescued into a transfectant virus if the cells are also infected with an influenza helper virus. Selection of the transfectant viruses can be performed by using neutralizing antibodies against the HA protein of the helper virus.
erated a recombinant influenza virus that expressed a CD8+ T-cell epitope derived from the circumsporozoite (CS) protein of Plasmodium yoelii in its HA. Mice immunized with this transfectant virus made a vigorous cytotoxic T lymphocyte response against this epitope (25). By boosting mice with a recombinant vaccinia virus expressing the CS protein, it was possible to achieve protective immunity (60%) against challenge with live P. yoelii sporozoites. Additional protective immune responses were generated by immunizing mice with transfectants expressing B-cell-specific epitopes located in the repeat region of the CS protein of P. yoelii. Up to 80% of immunized mice were immune to challenge with one hundred P. yoelii sporozoites (26).
Foreign epitopes can be inserted into several sites on the HA molecule of influenza viruses, and most conveniently into the stalk region of the NA. In fact, stretches of more than 80 foreign amino acids have been successfully inserted into the stalk region of the NA (27, 28) (S.Itamura, personal communication). Although some of these constructs show interesting biological properties, this approach of epitope grafting has its limitations in terms of the size and the nature of the epitope that can be expressed (since the chimeric protein may affect the viability of the recombinant virus).
A generic approach to the expression of foreign proteins is the construction of bicistronic genes which can be packaged into infectious particles. The foreign gene can replace the open reading frame of one of the influenza virus genes and the respective influenza virus protein is then translated from an internal ribosome entry site (IRES element) on the genetically engineered gene. Alternatively, the foreign protein can be translated from an internal IRES sequence. Expression of several foreign polypeptides was achieved in this way (16, 29). However, many constructs did not result in viable viruses (unpublished results). Attempts are currently being made to identify the factors which determine the limitations of this approach.
The second method for the expression of foreign proteins takes advantage of autoproteolytic elements placed within a fusion protein. For example, a virus was constructed that expresses a fusion protein consisting of the full-length chloramphenicol acetyltransferase (CAT) protein, the 2A protease of foot and mouth disease virus, and the viral NA (30). This virus was stably passaged and expressed copious amounts of CAT protein in infected cells. However, in all cases of the fusion protein constructs, the foreign protein contains a 16-amino acid extension derived from the 2A protease which may alter the biological properties of the foreign protein.
Rescue of Infectious Rabies Virus from cDNA
Like the segmented negative-strand RNA viruses, the Mononegavirales group packages its own RNA-dependent RNA polymerase into virus particles to initiate viral RNA synthesis. Thus, naked RNA alone is unable to drive the replication cycle. Several approaches were taken to rescue model and full-length RNAs. First, a Sendai virus-like RNA transcript was amplified and expressed by transfecting the naked model RNA into Sendai virus-infected cells (31). This experiment suggests that complementation in trans by the viral polymerase complex is required for the amplification and expression of the viral RNA-like reporter gene. Subsequently, in a remarkable study, Schnell et al. (32) succeeded in constructing a plasmid that expresses a full-length rabies virus RNA transcript from a T7 RNA polymerase promoter. The plasmid DNA containing this viral insert was transfected into cells infected with a recombinant vaccinia virus expressing the T7 polymerase. Three other plasmids expressing the rabies virus N, P and L proteins were also cotransfected into these cells. In this recombinant vaccinia virus-driven system, the presence of the viral polymerase complex and of a full-length viral RNA (in plus sense) led to the formation of recombinant rabies virus.
This system has been elegantly exploited to study the promoter elements of rabies virus RNA and to elucidate the interaction of this interesting virus with cells (33). Surprisingly, cells infected with a mutant lacking the virus’ only glycoprotein (G) were still able to bud from the cell surface, albeit at a 30-fold lower efficiency (34). This experiment revealed that the surface protein G exhibits an intrinsic exocytotic activity. The system was further developed to show that a hybrid G/HIV-1 glycoprotein was able to form pseudotypes with the “G-less” particle, thus changing the host range by restricting infection to CD4+ cells. This experiment clearly demonstrates that genetic engineering can redirect the host range and cell tropism of rabies viruses. This should prove helpful for the development of novel vaccines as well as for gene therapy.
Rescue of Other Nonsegmented Negative-Strand RNA Viruses
An effective DNA transfection system has also been developed for another rhabdovirus, vesicular stomatitis virus (VSV) (35, 36) (Fig. 3). Again, the polymerase complex (N, P, and L proteins) was expressed in cells from plasmids transcribed by a T7 RNA polymerase-containing vaccinia virus recombinant. Recombinant VSVs expressing an additional transcriptional unit were rescued and high-expression levels of heterologous proteins were achieved (37). In a dramatic experiment, the authors were able to construct a recombinant VSV expressing the CD4 protein. This protein was packaged at levels of up to 30% of the G protein itself, and the recombinant particle had an 18% greater length than wild-type virus due to the extra gene. These results illustrate that VSV is an effective vector to express foreign proteins at high levels, and that the virus is tolerant to the insertion of novel transcriptional units. Reverse genetics systems have also been developed for paramyxoviruses. In the case of measles virus, a cell line constitutively
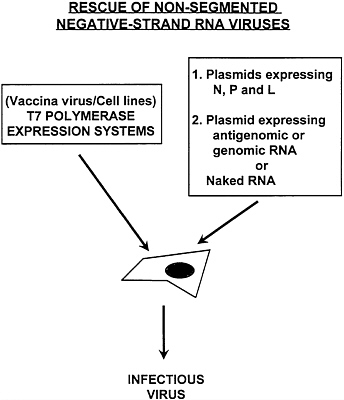
FIG. 3. Reverse genetics systems for the rescue of infectious nonsegmented negative-strand RNA viruses from cDNA. Transcriptionally competent viral RNPs are made by cellular expression of the viral proteins N, P, and L. This can be achieved by a variety of methods, including vaccinia virus-driven expression and/or complementing cell lines constitutively expressing T7 polymerase and viral proteins. The full-length viral RNA can be provided by transfecting plasmids expressing antigenomic or genomic RNA or by directly transfecting naked RNA (plus-sense or minus-sense). The intracellularly assembled RNPs are transcribed and replicated by the viral polymerase complex (N, P, and L proteins) generating infectious viruses.
expressing T7 polymerase and the measles N and P proteins has been used for the rescue of infectious virus from full-length clones (38) and vaccinia virus-based systems have allowed the rescue of respiratory syncytial virus (39) and of Sendai viruses (40, 41).
Most of the earlier systems developed for the nonsegmented viruses used the intracellular expression of antigenomic plus-sense RNA as the template to initiate the replication cycle. Either the plus-sense RNA was transcribed by the T7 polymerase expressed by a vaccinia-recombinant virus (32, 35, 36, 39–41), or transcription was driven by the T7 polymerase which was permanently expressed in cells (38). Recently, an important series of experiments showed that intracellular expression of a full-length transcript generated infectious Sendai virus regardless of whether the plus-sense or the minus-sense RNA was transcribed (41). Success appears to have come from fine tuning the system in terms of the concentration of the polymerase components (N, P, and L proteins) and from constructing plasmids giving rise to transcripts with 5′ and 3′ ends identical to those of the wild-type RNA. Optimization of the system also involved the use of the vaccinia virus inhibitors, cytosine arabinoside and rifampicin. These compounds reduced the cytotoxicity of vaccinia virus and resulted in a dramatic increase of the expression levels of a Sendai virus RNA-like reporter gene. Most interesting was the finding that recovery of infectious Sendai virus was also possible by transfecting naked RNA. The efficiency of recovery appeared to be lower using plus-sense RNA than the genomic minus-sense RNA (41). The latter results involving the use of naked RNAs extend the earlier findings that transfection of naked model RNAs alone results in the efficient amplification and expression of these minigenes in cells infected with Sendai virus (31), respiratory syncytial virus (42) or parainfluenza virus 3 (43, 44). In the future, improvements in the transfection systems to generate novel viruses with ease will provide us with even better tools for the study of negative-strand RNA viruses.
Perspective
The ability to genetically alter negative-strand RNA viruses has already enhanced this field of virology and may have a major influence on future developments in vaccines, gene therapy, cancer treatment, and manufacture of biologicals. First, structure-function studies of individual viral genes are now possible in the context of an infectious virus for a number of negative-strand RNA virus families. These groups consist of many medically important viruses including measles, mumps, respiratory syncytial, parainfluenza, influenza, and bunyaviruses. In the recent past, we tried to take a reductionist approach in virology; viral genes were studied in isolation by cloning and expressing them in different systems. The pendulum has now swung back in the other direction as we ask questions about how viral genes and gene products interact with host cell components and the host in general. This can best be done by studying genetically defined viruses and subjecting them to directed mutational analysis. These viral constructs are then available for biochemical analysis as well as for studying replication and growth in tissue culture or experimental animals. Obviously, structure-function studies of viral genes also include the analysis of promoter elements and other noncoding sequences.
Second, genetically engineered negative-strand RNA viruses should become candidates for use as live virus vaccines. Genetically engineered influenza viruses with changes in coding or noncoding sequences may induce immune responses which are longer-lasting and more protective than those generated by conventional influenza virus vaccines. In the case of respiratory syncytial and parainfluenza viruses, a recombinant DNA approach may be the only rational strategy, since the Jennerian approach has not resulted in acceptable vaccine candidates. Thus, tools are now available to design a new generation of vaccines for the medically important negative-strand RNA viruses.
Third, negative-strand RNA viruses may become useful vectors for the expression of foreign genes. Recombinant influenza viruses (16), rabies viruses (45), and VSV (37) have been used to express additional protein sequences or foreign genes. Packaging limitations and restrictions due to the length or the nature of the foreign gene are not yet defined for negative-strand RNA virus constructs, nor do we have sufficient information about the stability of these viruses once their genome structures have been extensively altered. These uncertainties notwithstanding, there is a major advantage in the use of negative strand RNA viruses as vectors (or as vaccines). These viruses do not go through a DNA phase and thus cannot transform cells by integrating their genetic information into the host cell genome. Furthermore, homologous recombination has never been observed for any of the negative-strand RNA viruses. Thus, replication-incompetent viral constructs grown in complementing cell lines should be free of contaminating virus generated by a recombinational event. In terms of safety, these properties weigh heavily in favor of negative-strand RNA virus vectors.
Novel viruses expressing foreign genes may serve prophylactically as vaccines, or they may play a role in gene therapy when a transient expression would be beneficial. The latter may be the case in cancer therapy, which could require a targeted infection followed by the induction of a lethal cytopathic effect. Repeated administration of negative-strand RNA viruses may not be feasible in this situation because of the host’s immune response. However, the choice of different
antigenic variants (as is possible with influenza viruses) may overcome this limitation.
Finally, the highly efficient expression of viral and foreign proteins via negative-strand RNA virus vectors may have additional biotechnological applications. It is possible that defective RNA constructs could be used for genetic immunization. This form of vaccination would resemble DNA vaccination (46) in that the defective particle would go through many replication rounds and persist without spreading to neighboring cells. Such RNA replicons may have interesting biological properties since the efficiency of infection should be comparable to that of whole viruses. Also, replication competent viral vectors may help in the manufacture of large quantities of biological reagents, since the quantities expressed by negative-sense RNA viruses can be high. It is also possible that purification of expressed proteins could be made easier if they were incorporated into extracellular virus particles.
The solutions to many of the issues discussed here will depend on the continuing success of basic science and the development of novel strategies to study viruses. Our horizons must expand and include the analysis not only of the viruses but also of their interactions with the host cell. Only by continuing to study these fundamental processes may we hope to reap the benefits offered to us by these new opportunities.
Work done in this laboratory was supported by National Institutes of Health grants to P.P.
1. Goff, S.P. & Berg, P. (1976) Cell 9, 695–705.
2. Post, L.E. & Roizman, B.R. (1981) Cell 25, 227–232.
3. Panicalli, D. & Paoletti, E. (1982) Proc. Natl. Acad. Sci. USA 79, 4927–4931.
4. Mackett, M., Smith, G.L. & Moss, B. (1982) Proc. Natl. Acad. Sci. USA 79, 7415–7419.
5. Jones, N. & Shenk, T. (1978) Cell 13, 181–188.
6. Samulski, R.J., Chang, L. & Shenk, T. (1989) J. Virol. 63, 3822–3828.
7. Cunningham, C. & Davison, A.J. (1993) Virology 197, 116–124.
8. Kemble, G., Duke, G., Winter, R., Spaete, R. & Mocarski, E.S. (1996) J. Virol. 70, 2044–2048.
9. Cohen, J.I., Wang, F., Mannick, J. & Kieff, E. (1989) Proc. Natl. Acad. Sci. USA 86, 9558–9562.
10. Wei, C.-M., Gibson, M., Spear, P.G. & Scolnick, E.M. (1981) J. Virol. 39, 935–944.
11. Mulligan, R.C. (1993) Science 260, 926–932.
12. Racaniello, V.R. & Baltimore, D. (1981) Science 214, 916–918.
13. Rice, C.M., Levis, R., Strauss, J.H. & Huang, H.V. (1987) J. Virol. 61, 3809–3819.
14. Liljestrom, P., Lusa, S., Huylebroeck, D. & Garoff, H. (1991) J. Virol. 65, 4107–4113.
15. Enami, M., Luytjes, W., Krystal, M. & Palese, P. (1990) Proc. Natl. Acad. Sci. USA 87, 3802–3805.
16. García-Sastre, A. & Palese, P. (1995) Biologicals 23, 171–178.
17. Subbarao, E.K., Kawaoka, Y. & Murphy, B.R. (1993) J. Virol. 67, 7223–7228.
18. Enami, M. & Palese, P. (1991) J. Virol. 65, 2711–2713.
19. Li, S., Xu, M. & Coelingh, K. (1995) Virus Res. 37, 153–161.
20. Yasuda, J., Bucher, D.J. & Ishihama, A. (1994) J. Virol. 68, 8141–8146.
21. Castrucci, M.R. & Kawaoka, Y. (1995) J. Virol. 69, 2725–2728.
22. Barclay, W.S. & Palese, P. (1995) J. Virol. 69, 1275–1279.
23. Pleschka, S., Jaskunas, S.R., Engelhardt, O.G., Zürcher, T., Palese, P. & García-Sastre, A. (1996) J. Virol. 70, 4188–4192.
24. Castrucci, M.R., Hou, S., Doherty, P.C. & Kawaoka, Y. (1994) J. Virol. 68, 3486–3490.
25. Li, S., Rodrigues, M., Rodriguez, D., Rodriguez, J.R., Esteban, M., Palese, P., Nussenzweig, R.S. & Zavala, F. (1993) Proc. Natl. Acad. Sci. USA 90, 5214–5218.
26. Rodrigues, M., Li, S., Murata, K., Rodriguez, D., Rodriguez, J.R., Bacik, I., Bennick, J.R., Yewdell, J.W., García-Sastre, A., Nussenzweig, R.S., Esteban, M., Palese, P. & Zavala, F. (1994) J. Immunol. 153, 4636–4648.
27. Castrucci, M.R. & Kawaoka, Y. (1993) J. Virol. 67, 759–764.
28. Luo, G., Chang, J. & Palese, P. (1993) Virus Res. 29, 141–153.
29. García-Sastre, A., Muster, T., Barclay, W.S., Percy, N. & Palese, P. (1994) J. Virol. 68, 6254–6261.
30. Percy, N., Barclay, W.S., García-Sastre, A. & Palese, P. (1994) J. Virol. 68, 4486–4492.
31. Park, K.H., Huang, T., Correia, F. & Krystal, M. (1991) Proc. Natl. Acad. Sci. USA 88, 5537–5541.
32. Schnell, M.J., Mebatsion, T. & Conzelmann, K.-K. (1994) EMBO J. 13, 4195–4203.
33. Mebatsion, T. & Conzelmann, K.-K. (1996) Proc. Natl. Acad. Sci. USA 93, 11366–11370.
34. Mebatsion, T., König, M. & Conzelmann, K.-K. (1996) Cell 84, 941–951.
35. Lawson, N.D., Stillman, E.A., Whitt, M.A. & Rose, J.K. (1995) Proc. Natl. Acad. Sci. USA 92, 4477–4481.
36. Whelan, S.P.J., Ball, L.A., Barr, J.N. & Wertz, G.T.W. (1995) Proc. Natl. Acad. Sci. USA 92, 8388–8392.
37. Schnell, M.J., Buonocore, L., Kretzschmar, E., Johnson, E. & Rose, J.K. (1996) Proc. Natl. Acad. Sci. USA 93, 11359–11365.
38. Radecke, F., Spielhofer, P., Schneider, H., Kaelin, K., Huber, M., Dotsch, C., Christiansen, G. & Billeter, M.A. (1995) EMBO J. 14, 5773–5784.
39. Collins, P.L., Hill, M.G., Camargo, E., Grosfeld, H., Chanock, R.M. & Murphy, B.R. (1995) Proc. Natl. Acad. Sci. USA 92, 11563–11567.
40. Garcin, D., Pelet, T., Calain, P., Roux, L., Curran, J. & Kolakofsky, D. (1995) EMBO J. 14, 6087–6094.
41. Kato, A., Sakai, Y., Shioda, T., Kondo, T., Nakanishi, M. & Nagai, Y. (1996) Genes Cells 1, 569–579.
42. Collins, P.L., Mink, M.A. & Stec, D.S. (1991) Proc. Natl. Acad. Sci. USA 88, 9663–9667.
43. De, B.P. & Banerjee, A.K. (1993) Virology 196, 344–348.
44. Dimock, K. & Collins, P.L. (1993) J. Virol. 67, 2772–2778.
45. Conzelmann, K.-K. (1996) J. Gen. Virol. 77, 381–389.
46. McClements, W.L., Armstrong, M.E., Keys, R.D. & Liu, M.A. (1996) Proc. Natl. Acad. Sci. USA 93, 11414–11420.





