4
Methods for Assessing Health Risks of Reclaimed Water
Any plan to augment potable water supplies with reclaimed water must include an evaluation of the potential health risks. Yet as described in earlier chapters, such assessment is complicated by several factors, including uncertainties about the potential contaminants and contaminant combinations that may be found in reclaimed water and about the human health effects those contaminants may cause.
This chapter discusses methods and strategies for assessing the health risks of drinking reclaimed water. Previous National Research Council (NRC) reports have provided similar guidance on assessing health risks of reclaimed water (see, most recently, Ground Water Recharge Using Waters of Impaired Quality, published in 1994). This chapter updates and expands on information in those earlier reports. The chapter also discusses the complications of and alternative strategies for using epidemiological studies to evaluate health risks of potable water reuse.
Evaluating Microbial Contaminants
Efforts to monitor water quality for microbiological safety have historically relied on measurements of one or more groups of coliform bacteria as indicators of fecal contamination, treatment efficiency, and the integrity of the water distribution systems. Fecal coliform bacteria are indicative of fecal contamination and associated health risks; however, the measurement and control of total coliforms (rather than only fecal coliforms) during disinfection is considered to be a more stringent treat-
ment goal. Water quality standards have used either or both measures depending on the type of water use.
While coliform bacteria serve well as indicators of bacterial pathogens, they do not predict the inactivation or removal of enteric protozoa and viruses (Gerba and Rose, 1990; LeChevallier and Norton, 1993; LeChevallier et al., 1991a, 1991b; Rose et al., 1991). For instance, LeChevallier and Norton (1993) found that multiple linear regression models using coliforms and temperature could predict only 57 percent of the variation in Giardia cyst concentration, whereas no model using indicator bacteria could adequately predict Cryptosporidium oocyst levels.
Methods to detect viruses in water were first developed by Paul and Trask (1947) for measuring enteroviruses in untreated wastewater. Coin (1966) was one of the first to detect viruses in finished drinking water meeting the existing coliform standard. In the 1970s, improvements in collection filters and the use of antibodies led to the first isolation of rotavirus and hepatitis A virus from water. Today, standard methods for the detection of enteric viruses are based on the ability of viable enteric viruses to destroy monkey kidney cells grown in vitro; this cell-destroying ability is known as cytopathic effect or CPE (Benenson, 1995).
Methods for detecting microbial protozoa were first developed for Entamoeba in the 1940s. Starting in 1965, research focused on the detection of Giardia. In the 1980s, a standardized approach for Giardia detection was developed that used filtration for collection and antibodies labeled with a fluorescent isothiocyanate (FITC) for enhanced microscopic detection (Rose et al., 1988a, 1988b). This approach was applied to the detection of Cryptosporidium after its first recorded waterborne outbreak in the United States in 1985.
As more protozoan and viral waterborne outbreaks occurred in waters meeting existing water quality standards, the limitations of using indicator bacteria became apparent. In response to these health concerns, the Environmental Protection Agency (EPA) promulgated the Surface Water Treatment Rule for drinking water in 1989 (U.S. EPA, 1989a, 1989b). The rule established treatment-based performance goals of 99.99 percent reductions of viruses and 99.9 percent reduction of Giardia. The rule also emphasized the use of sand or multimedia filtration for the removal of Giardia and the use of improved disinfection methods for the control of both viruses and Giardia. The target reduction level was based on anticipated levels of pathogens in ambient surface waters, and the performance goals were derived from a desired annual risk of microbial disease of not greater than 1 in 10,000. An Enhanced Surface Water Treatment Rule (U.S. EPA, 1996) is under development; the enhanced rule will include an assessment of Cryptosporidium in source waters and its removal by treatment processes.
Arizona is the only state that has standards for the concentration of enteric viruses and Giardia in reclaimed water; it is also the only state in which water reuse is regulated by a laboratory certification program, although specialized studies have been undertaken in Florida and California for viruses and protozoa.
To address the lack of information nationwide on levels of viral and protozoan pathogens and the efficacy of water treatment, EPA promulgated the Information Collection Rule (ICR) in 1996. Detection processes for the enteroviruses by cell culture and for protozoa by microscopy have been standardized for this rule, and laboratories are undergoing an approval process. The data will be used in future risk analyses to establish the necessary drinking water treatment performance criteria for the protection of public health. The results will be applicable to potable reclamation projects as well. However, when considering wastewater as a source of drinking water, particular attention should be paid to current limitations and other issues involved with the methodology used to detect microbial pathogens.
Microbial Detection Methods
Microbial detection methods can be described and compared in terms of recovery (the efficiency of the method for collecting microorganisms from water samples), sensitivity (a measure of the minimum number of microorganisms that can be detected per unit volume), and specificity (the proper taxonomic identification of the microbial agent). No method is 100 percent efficient; estimates of recovery tend to range from 5 to 60 percent. A method's sensitivity is often expressed as a detection limit, such as 1/100 ml, meaning that it is able to detect one microorganism in a 100 ml sample. In untreated wastewater, concentrations of microorganisms can be high enough to be readily detected in small test volumes. However, such methods are not sufficiently sensitive for testing the highly treated reclaimed water typically produced by potable reuse projects. With highly treated reclaimed water, larger volumes of water are needed for analysis, and microorganisms may occur at concentrations too low to be detected.
Table 4-1 presents the major microbial detection techniques as they are applied to the detection and quantification of bacteria, viruses, and protozoa. Culture techniques have long been used for the detection and enumeration of viable bacteria and viruses, while microscopy techniques have a long history in the identification of bacteria and protozoa.
Table 4-2 summarizes the advantages and disadvantages of some methods for evaluating the microbiological quality of reclaimed water. The polymerase chain reaction (PCR) has only recently been applied to
TABLE 4-1 Methods and Issues for Detection of Microorganisms in Water
TABLE 4-2 Advantages and Disadvantages of Microbiological Methods for Detecting Microorganisms in Reclaimed Water
|
Tool |
Advantages |
Disadvantages |
Application |
|
Culture of bacteria and viruses |
Widespread use |
In some cases, lacks specificity (i.e., generic for enteroviruses; HPC) |
To evaluate water quality and treatment, retrospectively |
|
|
Standardized |
|
For comparison to new methods and old databases |
|
|
Detects viable microorganisms |
Results take days or weeks |
When viability is of major concern |
|
|
|
Difficult to quantitate in some cases |
|
|
|
|
Measures only a fraction of the types of microorganisms present; no ability to detect those that are viable but nonculturable |
|
wastewater to identify a variety of microorganisms based on their species-specific nucleic acid sequence.
Use of Polymerase Chain Reaction Techniques
Polymerase chain reaction, or PCR, is a molecular technique used to detect a variety of microorganisms in environmental samples (Atlas et al., 1992; Bej et al., 1990; Johnson et al., 1995; Kopecka et al., 1993; Mahbubani et al., 1991; Tsai and Olson, 1991). PCR can rapidly identify a specific organism. However, before PCR can be used routinely for environmental monitoring, several issues must be addressed, including the test's sensitivity, the viability of detected pathogens, and assay interference by inhibitors in the spectrum.
The sensitivity (or the limit of detection) of PCR is constrained by the technology. In most cases, only very small volumes (100 µl or less) can be processed through the thermal cyclers (machines that control the sample temperatures during processing) used in PCR assays. Therefore, concentration of samples is necessary. (Alternatively, larger-capacity machines could be developed in order to increase the sample volume.)
Further, because PCR does not distinguish live from dead microorganisms, a cell cultivation procedure must be performed before the results have relevance to health risks. This is especially true for samples taken from water that has undergone disinfection. PCR may therefore be most useful in untreated waters (source waters, recreational waters, shellfish harvesting waters, ground waters) where viability can be assumed, or in the evaluation of processes designed to physically remove microbiological particles (such as membrane processes).
Water quality is also an issue, since physical and chemical constituents in water can mask the target nucleic acid or inhibit the enzyme reaction that the PCR process uses to amplify the target DNA, creating a false negative result. Recently developed antibody capture procedures appear to have great promise in addressing the problem of interference for both protozoa (Johnson et al., 1995) and viruses (Deng et al., 1994).
Finally, PCR remains only qualitative in that the results are presented as positive or negative. The development of quantitative techniques using PCR would be very useful for assessments of the microbiological quality of drinking water.
Detection of Bacteria
Several well-established methods for detecting and enumerating coliform and fecal coliform bacteria indicators exist and are useful for evaluating the effectiveness of disinfection in water and wastewater treat-
ment. Generally, any treatment process that inactivates the indicator bacteria also inactivates pathogenic enteric bacteria to similar degrees. Nevertheless, the use of indicator bacteria has limitations. For instance, recent findings suggest that very low levels (10 to 100 colony-forming units (CFU) per liter) of Salmonella may be related to incidences of reactive arthritis (Smith et al., 1993), suggesting that Salmonella should be measured directly. In addition, Singh and McFeters (1990) reported a noncultivable but viable state for Yersinia bacteria after disinfection. The public health significance of these noncultivable bacteria has not been fully assessed. Future regulations may dictate that greater reductions of pathogenic bacteria be achieved and documented.
Directly detecting pathogenic bacteria has traditionally been a tedious process, requiring many biochemical tests to identify the genus and/or species. Most testing has been done with presence/absence tests or the most probable number (MPN) approach. A standard MPN procedure has been developed for detecting Salmonella in sludge (U.S. EPA, 1992); however, the sensitivity of the test has been questioned (National Research Council, 1996). No standard procedure exists for Salmonella testing in reclaimed water. As described above, PCR is a rapid detection method that can be combined with more traditional cell culture techniques to assess viability of bacterial pathogens. This approach has been successfully applied in the food industry (Fung, 1994) and may hold promise for use in reclaimed water as well.
Detection of Protozoa
Detecting low concentrations of protozoan cysts or oocysts in highly treated reclaimed water requires passing large volumes of water through a filter with an appropriate pore size (typically 1.0 µm). Unfortunately, this method also concentrates unwanted constituents in the sampled water (e.g., particulates, precipitated minerals), and these constituents must then be separated in the subsequent analysis using a concentration and clarification procedure (LeChevallier and Trok, 1990; Rose et al., 1989). The semipurified sample, consisting of the larger organisms with some of the unwanted constituents removed, is then mixed with fluorescent-tagged monoclonal antibodies specific to the cyst and oocyst wall using an indirect fluorescent antibody (IFA) procedure. The sample can then be examined and the protozoa identified by one or more techniques, such as epifluorescence or differential interference contrast (DIC) microscopy (LeChevallier et al., 1991a).
The efficiency of cyst/oocyst recovery for Cryptosporidium and Giardia is quite variable, and losses occur at each of the various steps of the detection process; thus many of the organisms contained in the original
water sample will be lost. Reported efficiencies are: 88-99 percent for sample collection; 16-78 percent for filter elution; 66-77 percent for concentration and clarification; and 9-59 percent for microscopic detection using the IFA method (LeChevallier and Trok, 1990; LeChevallier et al., 1991a; Ongerth and Stibbs, 1987; Rose et al., 1988b, 1989, 1991b). Given these variations in recovery, current methods for detecting protozoa tend to underestimate the true concentrations in environmental samples.
Indirect fluorescent antibody techniques using antibodies labeled with fluorescent isothiocyanate have greatly enhanced the ability to detect Cryptosporidium and Giardia in environmental samples. However, most antibody techniques provide no species identification (e.g., bird versus mammalian isolates), nor do they determine whether cysts and oocysts are viable. LeChevallier et al. (1991a, 1991b) reported that 10 to 30 percent of the organisms detected by IFA were not empty and therefore viable. Other inaccuracies may occur due to background fluorescence from naturally fluorescing organisms and from nonspecific binding of the antibody. Such problems may produce either false positives or false negatives. Although false positives can create inconvenience or necessitate further testing, Clancy et al. (1994) noted that false negatives (failing to detect protozoa that are present) pose more serious problems.
A number of IFA systems have been developed for Cryptosporidium and Giardia (Garcia et al., 1987; Rose et al., 1989; Stibbs et al., 1988). New methods under development should allow a better assessment of enteric protozoa. These methods include the use of cell cultures to detect viable and infectious oocysts, the use of immunomagnetic separation (IMS) techniques to enhance the recovery of cysts/oocysts (Jakubowski et al., 1996; Linquist, 1997; Slifko et al., 1997), and the use of internal stains for improved identification and detection.
Detection of Viruses
Viruses may be concentrated from water using either electropositive or electronegative filters. Either can concentrate viruses from large volumes of water, but clogging, particularly in positively charged filters, may occur if the water is high in suspended solids or turbidity (Rose et al., 1989). The adsorbed viruses are eluted and concentrated to a smaller volume using an organic flocculation procedure. This final concentrate may be inoculated into cell cultures to detect cytopathic effects, or it may be tested by PCR or other types of tests.
Detection methods have primarily been developed for the enterovirus group, which consists of poliovirus, echoviruses, and coxsackieviruses. Microbiological studies in the San Diego potable reuse project demonstrated the feasibility of recovering 1 virus in 1000 liters of water.
When large volumes of water are sampled, one of the unresolved technical issues is whether a given filter will adsorb an equivalent amount of viruses per unit volume of water as more water is passed through the filter or whether the filter's performance will decrease as the quantity of water sampled increases. Further, some research has suggested that not all enteroviruses are adsorbed by filters at the same rate (Powelson and Gerba, 1995).
Cell culture techniques using CPE (cytopathic effect, or cell destruction) on monkey kidney cell lines provide the principal method for virus detection. In contrast to processing procedures for detecting protozoa, the debris collected along with viruses is more easily separated before cell culture. The types of viruses are further identified by the use of specific antibodies to neutralize the virus's cytopathic effect. However, these cell culture methods may detect only a small percentage of the viruses found in polluted waters because not all viruses will cause CPE. When Payment and Trudel (1987) used labeled antibodies to determine the numbers of infectious foci in the cell culture, they found levels of viruses up to 100 times greater than what they detected by CPE.
While enteroviruses are the most heavily studied waterborne viruses, the limited data on virus identification indicate that retroviruses typically occur at higher levels in wastewater (Gerba and Rose, 1990). Hurst et al. (1988) found that adenoviruses could be detected at levels 94 times higher than enteroviruses when using molecular techniques (gene probes) to verify the results of cell cultures.
In sum, although methods exist for detecting some viruses, we lack sufficient data to determine the concentrations and diversity of the many viruses of concern in wastewater, reclaimed water, and ambient waters (Hurst et al., 1989). These include emerging viruses of concern such as Norwalk viruses.
Indicator Techniques
Several types of microorganisms have been suggested as alternatives to coliform bacteria as indicators of water quality, fecal pollution, and public health risks. These include the bacteria Enterococcus and Clostridium perfringens and the F-specific coliphage bacterial virus.
The Enterococcus bacteria comprise a subgroup of the fecal Streptococci bacteria and include S. faecium, S. faecalis, S. durans, and related biotypes (Clausen et al., 1977). Enterococci are generally more resistant to water treatment than bacterial pathogens or fecal coliforms, and membrane filtration procedures are available for sampling them (Cohen and Shuval, 1973; Davies-Colley et al., 1994; Sinton et al., 1994). Some epidemiological investigations found that levels of Enterococcus correlated with
an increased incidence of gastrointestinal symptoms associated with exposure to polluted recreational waters (Cabelli et al., 1979; Dufour, 1984). The EPA has suggested that Enterococcus may be the best microbial indicator for ambient waters (U.S. EPA, 1986). A few states have adopted the use of Enterococcus, but most continue to use total and/or fecal coliforms for evaluating recreational water quality and the wastewater effluents that may impact these waters.
Clostridium perfringens is a pathogenic bacterium found in human and animal feces. Because it forms resistant spores, Clostridium has been recommended as a conservative indicator of water quality and as a valuable supplement to other water quality tests, particularly in situations where the detection of viruses or fecal contamination is desirable (Fujioka and Shizumura, 1985; Payment and Franco, 1993). This microorganism is consistently present in municipal wastewater at concentrations of 103 to 104 CFU/100 ml. Fairly rapid and simple membrane filtration methods are available for the enumeration of C. perfringens. Researchers have found a significant correlation between levels of C. perfringens and enteric viruses and protozoa in evaluations of the treatment efficiency of filtration and disinfection (Fujioka and Shizumura, 1985; Payment and Franco, 1993). These authors suggested that the removal of this indicator by a factor 7 to 8 log10 essentially ensures the removal of enteric protozoa and viruses at current limits of detection. However, no jurisdictions in the United States are known to have adopted Clostridium as a regulatory standard (Cabelli, 1997).
The coliphages are viruses that infect Escherichia coli, and therefore the presence of coliphages in water indicates the presence of their host E. coli, which is excreted by animals and humans. Coliphages may serve as better indicators for human enteric viruses than bacterial indicators do because coliphages more closely resemble human enteric viruses in size, shape, and resistance to treatment processes. In a comparison of untreated and treated wastewater, river water, treated river water, and treated lake water, Havelaar et al. (1993) found significant correlations between levels of coliphage and levels of enteric viruses. However, this correlation was not evident for the untreated and treated wastewater samples, which suggests that other unknown factors may complicate the use of this indicator when evaluating recent wastewater inputs into a water body.
Payment and Franco (1993) examined the removal of coliphages and enteroviruses in drinking water treatment. The total removal and/or inactivation of enteroviruses by the complete drinking water process was estimated at 7 log10, based on coliphage removal. Coliphages have also been used as biological tracers in the environment to evaluate the movement of septic tank effluent (Paul et al., 1995). An advantage of using
coliphages to evaluate treatment processes is that large concentrations can be seeded into pilot or full-scale systems, and low numbers can be monitored after dilution and reduction by treatment to indicate process removal (Rose et al., 1996). Such seeded studies may provide useful data more rapidly than monitoring of indigenous microorganisms can.
Microbial Risk Assessment
Human health risk assessment is a scientific process that attempts to identify and quantify the health risks from exposure to environmental hazards. The use of risk assessment in drinking water began in 1974 with the congressional mandates of the Safe Drinking Water Act. A series of studies and reports from the National Research Council evaluated health risks from contaminants in drinking water (National Research Council, 1977-1989). The EPA used the results of these studies and risk assessment to set maximum contaminant levels (MCLs) for a number of inorganic and organic chemicals. The Surface Water Treatment Rule (U.S. EPA, 1987) used a risk assessment approach, based on the presumed levels of pathogens in surface waters, to mandate certain treatment processes designed to reduce the annual risk of disease occurrence from viruses and protozoa to less than one disease occurrence per 10,000 people served (Regli et al., 1991).
Microbial risk assessments have been performed for nonpotable reclaimed water systems based on hypothetical exposure scenarios (Asano et al., 1992) and on monitoring data (Rose et al., 1996), but no approach has been formally developed for reclamation systems used for potable water supplies.
Recently, a conceptual framework for evaluating the risks associated with exposure to microbial contaminants has been developed (ILSI, 1996). This approach modifies a four-step paradigm developed by the National Research Council for chemical contaminants (see Box 4-1) into one better suited to microbial contaminants. The first phase of the process involves characterizing the human health effects and determining the dose-response relationship. This information comes from clinical, epidemiological, and public health surveillance data. The second phase involves estimating exposure and using the dose-response models to quantify the health risk. This two-phase approach is described below.
Human Health Effects and Dose-Response Modeling Microbial risk assessment requires adequate knowledge about the relationship between exposure to microorganisms and consequent health effects. Clinical and epidemiological studies provide the information nec-
|
Box 4-1 Steps in Risk Assessment In 1983 the National Research Council defined four steps in the risk assessment process:
|
essary to determine the dosage or exposure level necessary to cause infection or disease Infection is the colonization of microorganisms m the body and may or may not cause symptoms Infectious dose is often expressed as the dose required to cause an infection in 50 percent of the population who are exposed (ID50)
Infectious dose has been evaluated in human volunteers for many enteric microorganisms in order to develop dose-response curves Haas (1983) first developed dose-response models using data from human volunteers who ingested various levels of viruses, protozoan cysts or oocysts, or bacteria The individuals were then monitored for infection and, in some cases, disease The ratio of those infected to those exposed at each dose formed the basis for the dose-response curve
Some recently developed models provide a good fit to human doseresponse data sets Haas et al (1993) found that a beta-Poisson model best described the probability of virus infection, Rose et al (1991) and Haas et al (1996) also developed exponential models to evaluate the risks of Giardia and Cryptosporidum infections Table 4-3 summarizes the dose-response models for six different microorganisms from various studies Estimates of risk may be obtained by substituting a given value for exposure in the table, represented as N or number of microorganisms ingested
TABLE 4-3 Probability of Infection Models and Best Fit Dose-Response Parameters for Various Human Feeding Studies
|
Organism |
Best Modela |
Model Parametersb |
|
Rotavirus |
Beta-Poisson |
α = 0.26 β = 0.42 |
|
Giardia |
Exponential |
γ = 0.0198 |
|
Salmonella |
Exponential |
γ = 0.00752 |
|
E. coli |
Beta-Poisson |
α = 0.1705 β = 1.61 x 106 |
|
Shigella |
Beta-Poisson |
α = 0.248 β = 3.45 |
|
Cryptosporidium |
Exponential |
γ = 0.00467 |
|
a Models Pi = 1 - (1 + N/ß)-α (beta-Poisson) Pi = 1 - exp (-γN) (exponential) b Model parameters: Pi = probability of infection (ability of the organism to establish and reproduce in the intestine) N = exposure, expressed as numbers of microorganisms ingested (CFU of bacteria, cysts, or oocysts of Giardia or Cryptosporidium, or PFU (plaque-forming units) of viruses) α, β, γ = constants for specific organisms that define the dose-response model SOURCE: Haas et al., 1996; Rose et al., 1991a. |
||
Exposure and Risk Characterization
Using previously defined models for infection and data from human health effects, one can estimate the health risk a pathogen poses at various exposures. Table 4-4 compares four microbial contaminants for risks of infection, severity of symptoms, mortality in normal populations, and mortality in vulnerable populations. The values for exposure in drinking water were estimated based on concentrations reported in surface waters and using treatment reductions of 99.9 percent for Giardia, 99.99 percent for rotavirus, and 99.9999 percent for Salmonella. With these assumptions, the risk of infection ranged from a low of 2.7 x 10-4 to a high of 4.9 x 10-3, depending on the microorganism. While the dose models shown earlier (Table 4-3) indicate that the infectivities of rotavirus, Giardia, and Crypto-
TABLE 4-4 Comparative Risks of Disease and Mortality From Infections Acquired via Drinking Water
|
|
|
Daily Risk of Health Outcome |
|||
|
Etiologic Agent |
Daily Exposure (number of organisms)a |
Infectionb |
Severityc |
Mortalityd |
Vulnerable Mortalitye |
|
Rotavirus |
0.008 |
4.9 x 10-3 |
4.4 x 10-5 |
4.0 x 10-7 |
4.9 x 10-5 |
|
Salmonella |
0.116 |
2.7 x 10-4 |
1.1 x 10-5 |
2.7 x 10-7 |
1.0 x 10-5 |
|
Giardia |
0.132 |
2.6 x 10-3 |
1.8 x 10-5 |
2.6 x 10-9 |
|
|
Cryptosporidium |
0.96 |
4.5 x 10-3 |
4.1 x 10-5 |
4.1 x 10-9 |
2.1 x 10-5g |
|
a These are considered as high exposure estimates based on treatment reductions of 99.9% for Giardia, 99.99% for rotavirus, and 99.9999% for Salhmonella, and assuming consumption of 2 liters of water per day. b Daily risk of infection is based on the probability of infection modeled for the specific microorganism and the corresponding daily exposure given in the table. c Risk of severity is calculated as the number of individuals hospitalized, divided by the total number of cases for the particular microorganism that have been documented during waterborne outbreaks, and then multiplied by the probability of infection (taken from Rose et al., 1996). d General population. e Nursing homes. f No documented increase in mortality in the nursing home population. g Risk in immunocompromised. |
|||||
sporidium vary, their overall risks of infection are similar because of the differences in their potential exposure. The bacteria, represented by Salmonella, represent the lowest risk of infection. When evaluating mortality rather than just infection, the bacteria and viruses become more significant (by a hundredfold) than protozoa.
Risks from microbial contamination depend not only on the dose of microorganisms ingested but also on the host's immune status. Sensitive populations, including the elderly, infants, and people with compromised immune systems, stand at greater risk of severe outcomes (Gerba et al., 1996). For example, case-fatality ratios go from 0.001 percent for Cryptosporidium in the general population to 50 to 60 percent in the immunocompromised population (Rose, 1997). The immunocompromised population's risk of mortality is 100 times greater for viruses and bacteria and 10,000 times greater for Cryptosporidium.
An individual who contracts an infection from exposure to a waterborne agent may in turn serve as a source of infection for other individuals regardless of whether symptoms are apparent. These secondary infections should be considered when assessing the risk of a particular exposure.
Such secondary cases may arise by a variety of mechanisms. Among close family members, household secondary cases can arise by direct or indirect (such as from surface or food contamination) contact; this is particularly a concern when the infected individual is a child (Griffin and Tauxe, 1991; MacKenzie et al., 1995). Table 4-5 summarizes secondary transmission statistics obtained from a variety of waterborne and nonwaterborne outbreaks. It is unclear whether these secondary statistics would be applicable in nonoutbreak situations.
The spread of infectious disease in a population may be modeled using standard population-state models from mathematical epidemiology (Bailey, 1975, 1986). When applied to the transmission of waterborne infectious disease, such models contain many parameters, including some for which only poor or no estimates are available. However, a preliminary illustration of the methodology as applied to reclaimed potable water does exist (Eisenberg et al., 1996). These models account for typical numbers of infected, ill, recovering, and immunocompromised subpopulations in computing the passage of an infectious disease through a community.
Actual levels of exposure to microorganisms through drinking water have been difficult to estimate, because the data on reductions of many pathogens are limited and extrapolations are often needed to determine tap water concentrations. A limited amount of data on virus and protozoa removals during water treatment is available (LeChevallier et al., 1991a; Payment and Franco, 1993; Payment et al., 1985; Stetler et al., 1984).
TABLE 4-5 Summary of Secondary Case Data in Waterborne Disease Outbreaks
|
Organism |
Secondary Attack Ratioa |
Secondary Prevalence in Householdsb |
Source of Outbreak |
|
Cryptosporidium parvum |
0.33 |
0.33 |
Contaminated apple cider (Millard et al., 1994) |
|
C. parvum |
N/Ac |
0.042 |
Drinking water in Milwaukee (MacKenzie et al., 1995) |
|
Shigella |
0.28 |
0.26 |
Child day care center (Pickering et al., 1981) |
|
Rotavirus |
0.42 |
0.15 |
Child day care center (Pickering et al., 1981) |
|
Giardia lamblia |
1.33 |
0.11d |
Child day care center (Pickering et al., 1981) |
|
Viral gastroenteritis |
0.22 |
0.11d |
Drinking water in Colorado (Morens et al., 1979) |
|
Norwalk virus |
0.5-1.0 |
0.19 |
Swimming (Baron et al., 1982) |
|
Escherichia coli 0157:H7 |
N/A |
0.18d |
Child day care center (Spika et al., 1986) |
|
a Ratio of secondary cases to primary cases. b Proportion of households with one or more primary cases who have one or more secondary cases. c N/A = information not available. d Proportion of persons in contact with one or more primary cases who have a secondary case. |
|||
Nevertheless, the lack of monitoring data for evaluating exposure remains the greatest barrier to adequately assessing risks posed by microbial pathogens.
A final difficulty in assessing risks from microbial contamination of reclaimed water is in evaluating the environmental fate of pathogens in indirect potable reuse projects. As explained in Chapter 2, pathogens die slowly under certain environmental conditions, with the rate of die-off
dependent primarily on temperature. However, monitoring the fate of pathogens in the environment is extremely difficult due to dilution and a lack of knowledge of the hydrology of the receiving water.
These complicating factors introduce great uncertainty in the assessment of potential risks due to microbial contamination of reclaimed water. System reliability is therefore crucial. As discussed in Chapter 6, the reliability of treatment processes in reducing risks will need to be critically assessed, and multiple treatment barriers will be needed in case one of the processes fails. Further, public health officials involved with surveillance programs (also discussed in Chapter 6) in areas using reclaimed water will need to be aware of the symptoms associated with infections arising from waterborne pathogens, including emerging pathogens, and may need to broaden the scope of public health monitoring.
Evaluating the Risk of Chemical Contaminants
Evaluating the risk posed by chemical contaminants poses challenges in any context, and some unique challenges in reclaimed water.
Safety Evaluations of Reclaimed Water
Calculations of health risk from chemicals in drinking water are largely based on extrapolations of the results of toxicological experiments on animals and estimates of human exposure to the chemical. Based on such risk information, national standards have been established for a limited but growing number of chemical contaminants in drinking water. This approach has been considered adequate for drinking water derived from relatively pristine sources or from sources that have been used for a long time without evidence of harm. In this sense, the regulation of drinking water resembles the approach used for establishing the safety of commercially produced foods, in that it assumes the use of an approved (and relatively safe) source of raw materials, then concentrates on monitoring for common contaminants in these materials or for problems in their processing (e.g., a lack of good manufacturing practices).
However, current drinking water standards are not intended to ensure the safety of reclaimed water because the wastewater may introduce new, unknown, or unquantified sources of contamination (as explained in Chapter 2). For instance, we have poor toxicological data on the wide variety of organic compounds in wastewater. Thus, even if a water reuse system could identify all the organic material in its processed wastewater (which, as explained in Chapter 2, is impossible), there would be no basis for assigning risks to most of the identified compounds. Because of this
and because many compounds are unidentifiable, toxicological testing of reclaimed water may be the only way to ensure the water's safety. However, applying toxicological methods to reclaimed water is difficult because of the water's variable chemical composition and the difficulty of obtaining representative samples. The challenge becomes how to test high doses of essentially unknown contaminants so that standards with some margin of safety can be established.
The chemical hazards in reclaimed water derived from municipal wastewater may vary depending upon a number of factors, including (1) the composition of chemicals intentionally or unintentionally discharged to the system from domestic and industrial sources, (2) variability in the removal of these chemicals by treatment, (3) the introduction of chemicals during the treatment process, and (4) the creation of entirely new chemicals as a result of chemical reactions during the treatment. If all wastewaters had the same composition, one could extrapolate the apparent safe consumption of a drinking water augmented with treated wastewater from one community to another. However, the complexity and potential variability in the chemical composition of wastewaters from different localities make such extrapolation quite difficult.
Recent research on the toxicity of different chemical mixtures has shown that the main health hazards of a chemical mixture are frequently posed by very low concentrations of highly potent chemicals (National Institutes of Health, 1993). The potencies of chemicals as toxicants or as carcinogens easily span 6 to 9 orders of magnitude. Therefore, toxic effects of chemicals in a wastewater are not necessarily attributable to the chemicals that occur in the highest concentrations.
Another factor in evaluating uncharacterized mixtures from treated wastewater is the presence of large amounts of nontoxic chemicals that complicate the testing of mixtures. For example, most drinking water contains inorganic salts that pose absolutely no health concern at the concentrations at which they occur. Yet these salts must be separated out before the water's organic chemicals can be evaluated, because concentrating the salts would dehydrate and kill experimental animals in the same way that sea water would.
The earlier National Research Council review Quality Criteria for Water Reuse (NRC, 1982) made some recommendations related to the type of testing that should be applied to establishing the safety of reclaimed wastewater. Much of the remainder of this section assesses the adequacy of that approach. Of particular importance is the evolution of how the results of specific tests are interpreted, how they fit in an overall scheme of health-effect testing procedures, and how decision rules (sometimes unstated) are applied to the results of tests.

Checking the reverse osmosis unit at San Diego's Aqua III water reclamation facility.
Photo by Joe Klein.
Approaches to Toxicological Testing
The most common way to evaluate chemical hazards to human health is to test compounds in live animals (usually mammals), a process known as in vivo testing. For special purposes, testing of lower life forms, such as bacteria in cell cultures, is used; this is known as in vitro testing. Food and drug industries have developed fairly standardized testing strate-
gies of both types to detect adverse health effects of a given chemical with some level of certainty. First, screening tests identify which organs and/or physiological functions are most sensitive to the effects of the chemical. Then more specialized investigations assess the precise effects on physiological functions and seek to establish dose-response relationships. Conducted rigorously, such testing yields information that can be accurately extrapolated to humans.
In general, in vivo testing is the most effective screening method because mammals possess all the physiological systems that can be affected by a chemical in humans, and gross effects can be readily identified. In vitro investigations usually seek to evaluate the mechanism of a chemical's impact rather than its actual effects or, more recently, to provide direct sensitivity comparisons of human cells relative to those of the animals in which prior screening work has been done. Usually such information must be related back to the whole animal or human to determine how much active metabolite of a chemical reaches the receptor site at a given dose. Ideally, dose-response relationships are then developed to compare the internal concentration of the chemical resulting from external exposure of humans and experimental animals.
In vivo testing provides the best screening data but requires considerable investment. For example, experiments designed to detect the carcinogenic properties of a chemical generally involve treatments over a significant fraction of a test animal's life span. Reproductive or developmental effects require specific assessments of reproductive competence of sexually mature animals or observations of development in pregnant animals, which in turn requires observations over more than one generation.
Because of the expense involved in live animal testing, many in vitro tests have been developed for screening purposes with the hope that they would be predictive of carcinogenic and reproductive effects. These in vitro screening tests are not intended as a substitute for in vivo testing, but merely to identify chemicals needing further testing.
In the 1970s, a series of inexpensive in vitro tests was introduced into common use in safety testing. The most widely used was the Salmonella/ microsome assay, commonly known as the Ames test. Its purpose was to test for mutagenic activity, because mutation plays an important role in the development of cancer. This test was subsequently applied to a wide variety of environmental problems, including drinking water and reclaimed water, in the hopes that it would provide a cost-effective method for evaluating carcinogenic hazards in the environment. The apparent success of the test spawned an interest in developing in vitro techniques to detect other toxicological end points.
As illustrated in Table 4-6, a safety testing strategy emerged that in-
TABLE 4-6 Progression in Safety Testing
|
Screening/ Hazard Identification |
Dose-Response Determination |
Risk Assessment |
|
Short-term in vitro Point mutation Clastogenesis Reproductive effects Developmental effects Neurotoxicity Immunotoxicity Target organ testing |
Controlled human experimentation (in vivo can only be done in special circumstances, e.g., drug development) Animal studies, including two species tests for specific end points and studies of metabolism and pharmacokinetics |
Epidemiological studies Determination of human exposure Application of conventional risk assessment techniques to in vivo data |
|
Short-term in vivo Mutagenesis Clastogenesis Carcinogenesis Target organ identification Developmental effects Reproductive effects Nurotoxicity Immunotoxicity Endocrine disruption Metabolism effects |
|
|
corporates short-term in vitro and in vivo testing for screening purposes. Broadly speaking, the purpose of screening is to make sure that no significant health effect is overlooked. These testing schemes seek to establish a finite, cost-effective set of tests that can quickly and inexpensively identify the chemicals that pose enough possible risk to warrant more extensive testing. Depending the nature of the chemical, the extent to which prior toxicological data are available, and the chemical's intended use, a chemical may undergo testing by some or all of the screening tests listed in Table 4-6. For example, an organophosphorus pesticide (which works by interfering with neurological functions) would be tested both in vivo and in vitro for its ability to produce delayed neurotoxicity. If the screening tests suggest the compound has a toxic potential, then careful dose-response studies would be conducted. Further testing is especially important if a chemical seems capable of producing specific types of toxicity (e.g., neurotoxicity, reproductive toxicities, teratogenesis) at dose levels that are not lethal to the animal being tested. Generally the tests are run on at least two species to ensure that no large interspecies differences in responses are distorting the test results.
Conventionally, risk assessments are made from these data, with ap-
propriate uncertainty factors included to compensate for lack of direct data in humans. It is occasionally possible to make quantitative comparisons of internal dosimetry between experimental animals and humans if appropriate in vitro-to-in vivo and in vitro-to-in vitro comparisons of metabolism and pharmacokinetics of the chemical exist. These data, in turn, can be used to improve the accuracy of a conventional risk assessment procedure, which typically incorporates large uncertainty factors.
This general strategy has a history of use in the testing of food additives and new therapeutic drugs. The EPA has used the strategy under the Federal Insecticide, Fungicide, and Rodenticide Act and the Toxic Substances Control Act, and the Food and Drug Administration (FDA) uses the process as well. The general practice has been to test such products to the point where there is clear evidence of some toxic effect, then compare the toxic doses to the levels normally used in food or drugs to ensure an appropriate margin of safety.
The Role of Toxicological Testing in Water Reclamation Projects
The 1982 National Research Council evaluation of health considerations for the potable use of reclaimed water found that it was a practical impossibility to identify and test the toxicity of all of the individual compounds present in reused water, and that it was ultimately necessary to test the toxicity of mixtures of chemicals instead. Because the available toxicological tests were insensitive to the low concentrations present in reclaimed water, it was recommended that the mixtures be concentrated so that chemicals would be present in concentrations high enough to detect effects. Recognizing the complex and essentially unverifiable nature of the mixtures involved, the 1982 report recognized that the following factors complicate the experimental results of any toxicological testing:
- The changing consistency of samples can be dealt with only by frequent resampling and testing.
- Additive, synergistic, or antagonistic effects of the mixture components may vary with individual samples and change with time.
- The concentration procedures could influence the chemical or physical composition of samples.
- The chemical and physical stability of concentrates is essentially an unknown.
- The mechanics of sample preparation for administration to animals may influence results.
TABLE 4-7 Toxicological Tests Recommended by the 1982 NRC Report Quality Criteria for Water Reuse
|
Phase I |
Phase II |
Phase III |
|
In Vitro Mutagenicity In vitro transformation |
|
|
|
In Vivo Acute toxicity Teratogenicity Short-term repeated dose studies—14 days (includes cytogenetics assay) |
Subchronic 90-day study in at least one rodent species, preferably in two species Reproductive toxicity |
Chronic lifetime feeding study in one species of rodent |
|
SOURCE: NRC, 1982. |
||
Notwithstanding these complications, and based on the general testing logic outlined in Table 4-6, the 1982 NRC panel recommended that a final comparison should be made between reused and conventional water based on the outcomes of a series of tiered tests designed to give information on the relative toxicities of the concentrates from the two water supplies (Table 4-7). Phase I of this recommended protocol includes in vitro assessments of mutagenic and carcinogenic potential by means of microbial and mammalian cell mutation and in vivo evaluations of acute and short-term subchronic toxicity, teratogenicity, and clastogenicity. Phase II includes a longer term (90-day) subchronic study and a test for reproductive toxicity on live animals. Phase III is a chronic lifetime animal feeding study.
Several studies of reclaimed water have included toxicological testing of organic concentrates derived from water. Unfortunately, in most of these studies the selected screening-level tests that correspond to NRC's Phase I were not always followed up with the more detailed Phase II and III studies necessary to confirm and elaborate on the results of the screening tests. As shown in Chapter 5, much of the testing was limited to in vitro tests designed to detect specific end points (e.g., mutagenicity and cytotoxicity). The value of in vitro testing is unclear when used alone to examine low-level risks in the environment. Much of the problem arises from inadequate follow-through on the decision rules of safety evaluations. These issues are explored in more detail in the following discussion.
Limitations of In Vitro Testing
There are several reasons why in vitro testing alone does not reliably measure health risk. As used in product safety testing, the original vision for mutagenicity tests was that they would have a low false negative rate for carcinogenic compounds and that any false positive data would be revealed by further testing. In practical terms, if a test is positive, a decision can be made either to drop the product altogether or to test further with systems that better assess the chemical's carcinogenicity.
However, several studies have shown the Ames test to be a poor predictor of carcinogenicity in general (Ashby and Tennant, 1988; Douglas et al., 1988; Parodi et al., 1983). A particular concern is a higher than anticipated false negative rate in these tests that has not been improved significantly by combining different types of tests for mutagenic effects.
To be useful (i.e., to provide information that cannot be gained in another way), in vitro test systems must focus on a single end point (e.g., carcinogenicity, mutagenicity, teratogenicity, reproductive toxicity, liver toxicity, neurotoxicity, kidney toxicity) or, in some cases, on one of several modes of action that can produce an end point (e.g., on mutagenesis, which can produce cancer). No single in vitro testing system addresses a substantial fraction of the many potential health effects that chemicals can have on animals or humans, and there seems little hope that any combination of in vitro tests will do so or be able to substitute for experiments on live animals in the foreseeable future. Negative results in a single mutagenesis test, for instance, provide no confidence that the chemical or product causes no nerve damage, liver damage, reproductive problems, and so on.
In vitro test systems do have considerable value as tools to resolve questions of extrapolation, for they allow very directed studies of mechanisms of action. However, such studies assume that some specific type of pathology has already been identified in an intact animal or human. Therefore, the toxicological use of in vitro systems is moving away from screening toward risk assessment of identified contaminants. The results of such risk assessment studies are used to select the extrapolation model rather than to provide data that would be used in a quantitative fashion.
These limitations raise the question of whether screening tests for specific end points should involve in vitro tests at all. Only in intact animals can all forms of toxicity be recognized in one test system. This becomes even more important when the material being tested is a complex mixture such as reclaimed wastewater. Using a large battery of tests to identify each and every potential toxic effect of a complex chemical composition that varies in time and place would be very expensive and time-consuming and produce results of questionable utility. Moreover,
testing must eventually return to an in vivo system before in vitro data can be related in any plausible way to actual risk.
Extrapolation of Test Results to Human Risk
Toxicological investigations of food additives or drugs make extrapolations from experimental systems to the human system and from the high experimental doses to the lower doses that are usually encountered by humans. Extrapolation from the high doses used in toxicological experiments to the low doses that might be present in reclaimed water creates uncertainty because effects that occur at high doses may not occur at low doses. In the case of presumed stochastic responses, particularly those that arise from chemically induced mutation, an assumption is sometimes made that low doses create the same responses as high doses, but with less frequency or severity—that is, that the response curve is linear. But this assumption ignores the role of nonmutagenic processes that occur only at high doses, such as those used in studies in experimental animals. These problems can be addressed by research on individual chemicals. However, the data that would allow reasonable quantitative conclusions to be made are still quite limited and do not clearly apply to the highly complex mixtures of chemicals found in wastewaters.
For these reasons and others, a recent workshop on molecular and cellular approaches to risk assessment concluded that the current state of knowledge does not allow meaningful risk assessments to be made purely on in vitro data (Sutter, 1995). Workshop participants recommended parallel in vitro/in vivo experiments in animals to validate an in vitro-to-in vivo extrapolation. In vitro measurements should then be made for critical end points in the appropriate human cells before the results can be extrapolated to humans (see Figure 4-1).
When considering potable reuse of treated wastewater, extrapolations to other locations and forward through time may also be necessary. While the few fairly extensive toxicological studies of reclaimed water that have been conducted (see Chapter 5) have shown no toxicological effects produced by reclaimed water, the extrapolation of these results to other reuse systems is greatly complicated by possible differences between municipal wastewaters from different systems, potential changes in wastewater composition over time, and the types of monitoring that would be needed to ensure continued toxicological safety of the reclaimed water.
Further complicating these extrapolations are the many chemicals present at low dose levels in drinking water from either natural or wastewater-influenced sources, which may interact with one another in an additive, synergistic, or antagonistic manner to produce biological effects
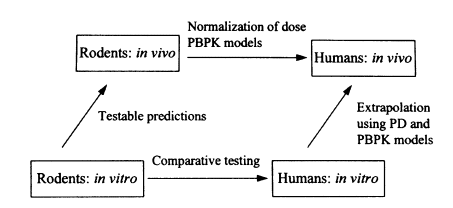
FIGURE 4-1 Parallelogram approach to risk assessment from whole-animal and in vitro testing procedures. The essential features are to confirm the critical toxicological response at the in vitro level and to adjust predictions of risk for humans based upon confirmation of in vivo observations in the appropriate target cell in intact animals. Development of the dose-response curve for humans would then involve modeling of target site dose with physiologically based pharmacokinetic (PBPK) models and response by pharmacodynamic (PD) models.
(NRC, 1989). Further, individuals within an exposed population vary in their susceptibility to chemical-induced injury, depending on factors such as age, nutritional status, gender, or genetic predisposition. Immunocompromised individuals, such as those undergoing organ transplant treatments or cancer chemotherapy or those infected with HIV, may suffer further immunosuppressive effects by exposure to some chemicals in drinking water at even very low dose levels. Given the millions of people using public drinking water supplies, it is reasonable to expect that such scenarios currently exist, and they should be factored into risk assessment strategies for treatment of public water supplies from any origin.
There is often limited information regarding how closely a sample used for testing represents the original constituents (see Chapter 2). Further, there has been little follow-up on the relationship between in vitro bioassays and live animal studies using samples derived from reclaimed water concentrates. Only two studies—one in Denver (Lauer et al., 1990) and one in Tampa (CH2M Hill, 1993)—have used live animal testing. While the Tampa study documented mutagenic activity in some samples through in vitro testing, no sample produced a significant positive response in vivo. Therefore, within the limits of the sensitivity of the in vivo tests that were used, the mutagenicity tests were not predictive.
In evaluating the safety of reclaimed water, additional uncertainties
also exist in extrapolating the results from one location to another and from the present to the future at a given location. The confidence with which data can be extrapolated from one circumstance to another depends upon the completeness of the data and the representativeness of the samples.
In setting guidelines for evaluating the chemical safety of reclaimed water, it may be useful to consider the two general approaches used in safety testing of other products. The first approach is used for products whose chemical composition can be reasonably well defined. In general, such products are pure chemicals or relatively simple mixtures. For such products, very specific safety standards can be established.
A second approach, for products having poorly defined chemical composition, is to provide strict guidelines on production procedures. Standards on the chemical composition of such products can still apply, but they are supplemented by the guidelines and regulations regarding production techniques. For this type of situation, manufacturing may be approved on a plant-by-plant basis to ensure that production methods meet relevant guidelines.
Both approaches are used to regulate drinking water from non-reclaimed sources: maximum contaminant levels set explicit standards for specific chemicals, but treatment processes are also regulated. A similar combined approach can be used to regulate the toxicological safety of reclaimed water. That is, standards can be set specifying the types of toxicological testing that the water must undergo, but the processes for ''manufacturing" the reclaimed water can and should also be strictly regulated.
Epidemiological Methods for Evaluating Health Risks
Human epidemiologic studies attempt to measure adverse health effects in a population exposed to a health hazard. A related but distinct activity is public health surveillance (described in Chapter 6), which collects information on morbidity and mortality but does not determine risk. Surveillance systems may reveal trends in disease morbidity and mortality but do not necessarily relate these trends to water quality. Determining whether an observed pattern of disease is associated with exposures to contaminants in drinking water requires a specifically designed epidemiologic study.
The use of epidemiologic methods to study health risks associated with drinking water has been recently reviewed by Craun et al. (1996) and Savitz and Moe (1997). Most epidemiologic investigations of drinking water and health have been conducted following waterborne disease
outbreaks, when the investigators were primarily concerned with identifying the factors causing the particular outbreak and controlling the outbreak as quickly as possible. Epidemiologic studies designed to detect endemic waterborne disease or other health risks that may be associated with low levels of microbial or chemical contaminants in drinking water have proved more challenging.
In situations where there have been no obvious health problems or observed outbreaks and where endemic waterborne disease is low, the study population must be large enough to allow investigators to accurately measure and calculate whether a true difference exists between the disease patterns in the exposed population and the disease patterns in the unexposed population. In the study design phase, statistical-power calculations can be used to indicate the minimum risk that can be detected with a specific study population size. In the data-analysis phase, statistical-power calculations can also be used to interpret negative findings by providing information about the minimum risk that could have been detected given the size of the study population and the measured disease rates. Epidemiologic studies cannot prove that reclaimed water poses no risk; rather, negative findings can only show that the risk, if any, was less than what the study was capable of detecting.
Another critical issue in epidemiologic studies of reclaimed water is the problem of comparability. Any epidemiologic study requires comparisons of health outcomes or exposure experiences between different populations. In areas using or considering the use of reclaimed water, reclaimed water may be of better quality than other area water sources. If so, defining the appropriate "unexposed" or control group is difficult. The "unexposed" group may be a population in the same geographic area that consumes poor-quality water that is not reclaimed water, a population in a different geographic area that consumes high-quality water that is not available in the study area, or a population consuming bottled water. When considering the possible health risks associated with potable water reuse, one must also consider the risks associated with these other available water sources.
Basic Study Designs
Ecologic studies, also known as geographic studies, are often used in investigating the possible health effects of exposure to environmental contaminants. These studies are descriptive, are relatively inexpensive and easy to perform, and usually take advantage of existing data on mortality, morbidity, and demographics. Exposure and health outcome are characterized on an aggregate level. For example, initial studies of the potential health risks from chlorination by-products in drinking water
used ecologic designs in which exposure and cancer rates were characterized on a community level. Cancer mortality rates in communities with chlorinated surface water supplies were compared to the rates in control or "unexposed" communities served by ground water.
Ecologic studies are valuable for developing hypotheses that can be tested in analytical epidemiologic studies. However, ecologic studies have limitations. For instance, such studies do not take into account individual risk factors (such as smoking or occupational exposure) or individual water exposure patterns (such as the use of bottled water or the length of residence in the study community). In addition, they assume that the distribution of individual risk factors and behaviors will be relatively equal in the exposed and control communities. In studies examining potential health effects with long latency periods (such as cancer), the results may be weakened by a mobile study population, since migration in and out of the study area over time will reduce the sample size of the truly exposed population.
In another type of study, the case-control study, the exposure histories of individuals with the disease of interest ("cases") are compared to the exposure of individuals without the disease ("controls"). With this design, cases and controls are queried directly about their residence history and water consumption habits, which provides greatly improved estimates of risk associated with exposure to water from different sources. Case-control studies allow the association between exposure and a single disease or health outcome to be evaluated while controlling for individual risk factors. However, case-control studies cannot prove that the exposure caused the disease or health outcome, because they do not provide evidence that the exposure preceded the disease. Case-control studies are useful in examining risk factors for specific health outcomes and are generally more efficient than cohort studies (described below), especially for rare health outcomes, because they require fewer participants for adequate statistical power.
In a cohort study, also known as a longitudinal study, the disease rates among a group of people who are exposed to the substance of interest (such as reclaimed water) are compared over time to the disease rates in a group of people who are not exposed to the substance. Because this design identifies the study population and determines exposure before the development of disease, it can be used to determine the temporal relationship between exposure to reclaimed water and the development of various health outcomes. These studies are typically the most expensive epidemiologic designs, especially if they require long follow-up times. Retrospective cohort studies cost less and take less time than prospective ones. Studies of occupational exposures have used retrospective cohort designs in situations where historical exposure of a group of indi-
viduals ("cohort") was well characterized; health outcomes are then measured in the past or present. Although this study design has some limitations (such as the difficulty of collecting accurate historical exposure data), it has potential for future epidemiologic studies of possible health effects associated with potable reuse.
Each of these study designs has strengths and limitations, and no single study can conclusively establish the risks that may be associated with exposure to reclaimed water. Epidemiologists must examine a body of evidence collected from several studies and study designs and consider the following questions to determine whether the association between exposure and disease is a causal relationship:
- Temporal association: Did the exposure of concern precede the development of disease?
- Study precision and validity: Was the study well designed and conducted, and was the study population sufficiently large to detect meaningful differences in exposure and health outcomes?
- Strength of association: Was the measure of association large enough to be a credible rather than a spurious result?
- Consistency: Is there consistency of results across several studies with various designs and study conditions?
- Specificity: Does the exposure result in a specific health outcome?
- Biological plausibility: Is there scientific evidence from other fields (such as toxicology) to suggest that the exposure of concern results in the observed health effects?
- Dose-response relationship: Was there evidence that increased exposure resulted in greater risk and/or more severe health outcomes?
- Reversibility: Does removal of the exposure of interest result in a reduction of risk and/or disease?
-
The absence of one or more of the above conditions does not rule out causality. For example, some causal relationships can be weak and can result in multiple health outcomes. However, taken together, these criteria allow critical evaluation of specific epidemiologic findings in a broader context.
Exposure Assessment
Estimating actual exposure to microbial pathogens or chemical contaminants in drinking water is a difficult task. In general, populations who are served by a given water supply are considered to be "exposed." Yet there can be a significant range in the degree and mode of exposure, depending on individual water consumption habits, household treatment
devices, and variabilities within the treatment and distribution system. In addition, inhalation and contact also serve as potential transmission routes for waterborne diseases and must be included in estimates of exposure (Savitz and Moe, 1997).
In epidemiologic practice, exposure to waterborne chemical or infectious agents is assessed by either (1) routinely measuring the actual microbial pathogen or chemical contaminant of concern or a proxy (such as a microbial indicator organism) in tap water or (2) estimating the presence, and possibly the amount, of the contaminant based on characteristics of the water source and treatment processes (such as a chlorinated surface water supply versus an untreated ground water supply). Individual consumption of or exposure to the water supply can then be determined by conducting interviews that collect information on lifetime residence history and self-reported water consumption habits (Savitz and Moe, 1997). Attempts to reconstruct the lifetime water exposure history of deceased cases by interviewing family members or using the last residence listed on death certificates should be avoided since these methods may misclassify exposure and introduce bias into the study (Craun et al., 1996).
Biomarkers of Exposure or Susceptibility
There has been increasing interest in the use of biomarkers (biochemical or molecular markers) of exposure or susceptibility in epidemiologic studies. For example, epidemiologists can use serological surveys to determine the prevalence of specific antibodies in a population and then compare the prevalence of specific infections, such as Cryptosporidium, between populations with different water sources in order to study the endemic risks of waterborne cryptosporidiosis (Craun et al., 1996). Such techniques may strengthen our ability to detect and classify exposure and disease (especially early stages of disease), clarify the steps between exposure and development of disease, and study the role of host factors that may account for variation in response. These techniques also may enhance the use of epidemiologic data to provide individual and group risk assessments (Schulte, 1993). Biomarkers could be particularly valuable for studying health effects of reclaimed water where there are a variety of possible exposures (microbial and chemical) and health outcomes of interest (infectious diseases, cancers).
Biomarkers in epidemiologic studies should be approached with caution since they can introduce confounding factors or bias (Pearce et al., 1995). For example, appropriate laboratory techniques must be chosen to measure the relevant antibodies, and the results must be interpreted carefully, since the presence of antibodies to a particular microorganism does
not explain whether the infection was water related and may not indicate how long ago the infection occurred.
Sensitivities to chemical agents can also be highly variable. Genetic polymorphisms are proteins of the same general class that exist in different forms. These different forms have different functional properties. Individuals who express one form versus another can exhibit different sensitivities to chemical or microbial agents. As a consequence, molecular markers of genetic polymorphisms can be used to identify susceptible subpopulations and can enhance epidemiologic research by more accurately describing risk in specific groups, rather than assuming an average risk for the whole population. For instance, a biomarker that may be of relevance to health-effect studies of reclaimed water is the glutathione Stransferase theta 1 (GSTT1) gene. It has been hypothesized that polymorphisms in the GSTT1 gene may contribute to susceptibility to colon and rectal cancer. GSTT1 is involved in detoxification of a variety of halogenated organic compounds, including organic compounds present in chlorinated drinking water (bromoform, dibromochloromethane, dichlorobromomethane, and dichloromethane) (Casanova et al., 1997; Smith et al., 1995; Thier et al., 1993). Laboratory studies suggest that individuals with one or more intact copies of the GSTT1 gene could experience a different risk for colon and rectal cancer if exposed to chlorination byproducts or other halogenated organic compounds in water.
Outcome Measurement
As discussed in Chapter 3, microbial agents in water have been associated with a diverse range of health problems. Waterborne enteric and aquatic microorganisms can cause acute and persistent gastroenteritis, dysentery, infectious hepatitis, febrile illness, meningitis, myocardiopathy, herpangina, encephalitis, hemolytic uremic syndrome, poliomyelitis, conjunctivitis, pharyngitis, ear infections, wound infections, dermatitis, Legionnaires' disease, Pontiac fever, stomach cancer, and various types of toxin poisoning. Chemical agents in water are known to have various acute and chronic toxic effects, and to cause reproductive problems, and they have been associated with cancer (bladder, liver, colon, rectal, kidney, esophagus, stomach, pancreas, and breast), depending on the type of agent, concentration, and duration and route of exposure.
Waterborne illnesses range from common, mild symptoms to rare, severe events, with results ranging from a successful natural immune response to death (see Chapter 3). Given this wide array of health effects and the variety of types and concentrations of contaminants that may be found in reclaimed water, it can be difficult to choose appropriate health outcomes for epidemiologic studies of reclaimed water. The choice of
health outcomes to focus on for a particular study will affect the size of the study population and choice of study design. Some designs, such as prospective cohort and cross-sectional, are more amenable to the examination of frequent episodes of common illnesses, while other designs (case-control and retrospective cohort) are recommended for the study of rare, severe outcomes. Measuring both acute and chronic health outcomes in a single study is very challenging, because different measurement strategies may be required. Changes in disease incidence are a more sensitive outcome measure for environmental exposure than death, because (1) incidence would be expected to change earlier than mortality in response to an environmental exposure, (2) incidence is not affected by survival factors, and (3) incidence data are gathered from diagnostic information in patient medical records, which may be more informative than cause-of-death information from death certificates (Sloss et al., 1996).
The best method for measuring health outcomes in epidemiologic studies of health effects associated with water exposure depends on the study design and the health outcomes in question. Prospective studies of endemic waterborne infectious diseases have used personal health diaries to record episodes of gastrointestinal symptoms (Payment et al., 1991). Case-control studies of disinfection by-products and cancer have used medical records or death certificates (Craun, 1993). Ecologic studies of reclaimed water have used passive surveillance for infectious diseases and tumor registries and death certificates for evidence of chronic diseases (Sloss et al., 1996). Some outbreak investigations have used various forms of active surveillance, including lab-based active surveillance, telephone surveys, and monitoring of illness in institutions such as nursing homes and other "sentinel" or high-risk populations (MacKenzie et al., 1994). The quality of health outcome data collected by these diverse methods can vary, and data quality should be considered in the interpretation of results. Underdiagnosis and underreporting of disease and misclassification of health status may bias the results of an epidemiologic study.
Epidemiologic studies of water-associated health effects must consider several attributes of the exposure-disease relationships of interest. The timing of exposure relative to development and detection of disease varies considerably for different health outcomes. For most chemical agents suspected of causing cancer, risk arises from prolonged exposure, and there may be prolonged latency. Epidemiologic studies of these health outcomes must choose study populations who have had (or will have) sufficient lengths of exposure and follow-up time. For some cancers the latency time may be as long as 15 to 30 years. In contrast, chemical exposure that may affect reproductive health outcomes is of interest only during a brief interval early in pregnancy. Waterborne infectious
diseases typically have short incubation periods and occur in response to a "slug" of microbial contamination in the water supply. For both chronic and acute diseases, the time course can vary depending on dose of the agent, prior exposure to the agent, and specific host factors (Savitz and Moe, 1997).
The ability to attribute a specific health outcome to a particular waterborne agent is related to the specificity of the exposure-disease relationship (Savitz and Moe, 1997). Diseases such as acute gastroenteritis and bladder cancer have many known and potential causes, some of which are completely unrelated to waterborne exposures. Of all reported cases of a particular disease or set of symptoms, only a subset can be potentially attributable to specific water exposures. For infectious diseases, clusters of cases due to a point source of contamination may be identified by molecular typing of clinical and environmental isolates. In contrast, it is extremely difficult to ascribe chronic diseases such as cancer to specific waterborne contaminants. When there are multiple transmission routes of the contaminant or multiple causes of the health outcome, the study must consider the effects of confounding factors (such as occupational exposures, smoking, and health behaviors) that can distort the relationship between water exposure and health outcome.
Sources of Bias and Error
When planning epidemiologic studies or reviewing the results of previous studies, one must examine whether the results could be affected by systematic errors in the sampling strategy or data collection procedures. Nonrandom error in a study that leads to distorted results is called "bias." Epidemiologists have described four major types of bias (Greenberg, 1993), and certain types of study designs are more prone to specific types of biases.
Usually, the first consideration is the potential for bias in the selection of the study subjects, or selection bias. The study must select subjects who are representative of the population of interest. Selection bias is a design issue and cannot be corrected in data analysis. In ecologic studies, the whole community is theoretically included in the study. However, certain segments of the population may be underrepresented because of the way health outcome data are collected. For example, health care facilities in poor neighborhoods may not have the resources to accurately diagnose and report specific health outcomes. When selecting control communities for ecologic studies, it is important for the control community to be similar to the test community in every way (demographics, income, education, etc.) except for exposure to the substance of interest (i.e., the water supply). In case-control studies, how cases and
controls are selected is critical to the correct design of the study. Cases and controls must be selected without knowledge of their exposure. The strengths and limitations of various recruitment strategies need to be carefully considered. For example, choosing prevalent cases of cancer rather than incident cases will favor survivors; choosing controls from a hospital may select people with disease conditions that could also be influenced by environmental factors; and choosing community controls via telephone recruitment will favor those who have telephones and are available to answer the telephone (i.e., those who spend more time in the home). In cohort studies, the study population must be selected without knowledge of disease. Loss of certain members of the cohort during the follow-up period may result in bias if those who drop out of the cohort are unique in a certain way and become underrepresented in the final cohort. An example of such a unique group is families with young children; these families may find that they do not have time to participate in studies that require maintaining health diaries or responding to extensive interviews.
Information or misclassification bias can occur in the determination of exposure or of health outcome. This bias results from systematic errors in measuring either the exposure or the disease. In water studies, misclassification of exposure is a major concern. Information on an individual's exposure to a specific water supply may be based on information from death certificates or water utility bills. However, this measure only indicates exposure at a specific point in time that may not be the exposure time frame relevant to the disease of interest. For example, risks of bladder cancer may be strongly associated with the type of water supply an individual was exposed to 20 years previously and may not be related to that person's current water supply. When evaluating this type of bias, it is important to determine whether it is "differential" or "nondifferential." Differential misclassification occurs when either (1) exposure was measured differently for cases than for noncases or (2) health outcomes were measured differently for exposed than for unexposed people. Differential misclassification will result in either an overestimate or an underestimate of the measure of association between exposure and disease. Nondifferential misclassification occurs when either (1) exposure is measured equally incorrectly for both cases and controls or (2) health outcomes are determined equally incorrectly for the exposed and unexposed people. In these situations, the measure of association between exposure and disease is underestimated.
Recall bias can be a concern when data are gathered retrospectively and is most likely to occur in exposure assessment. For example, lifetime water histories may be recalled differently by cases than by controls, be-
cause ill people may be more likely to remember (and blame) specific exposures for their illness.
Confounding bias is a problem when the exposed and unexposed groups differ in the occurrence of some factor or factors that affect the development of the health outcome of interest. Common confounders are demographic characteristics such as age, gender, race, and occupation or behavioral characteristics such as smoking or alcohol abuse. For example, in an ecologic study, the study community may house a chemical manufacturing plant, and therefore it may have more individuals with occupational exposure to hazardous chemicals than the control community. This difference, rather than water supply, may account for higher cancer rates in the study community. The effect of confounding bias on the measure of association can be controlled in the study design either by restricting the study population to persons with a narrow range of a confounder (such as white, male, nonsmokers between the ages of 30 and 50) or by matching study groups based on confounders (such as matching cases and controls for age, gender, race); or confounding bias can be controlled in the analysis phase of the study by using techniques that account for the effect of the bias. All of these control methods rely on being able to identify and measure the relevant confounders for the health outcome of interest.
Conclusions and Recommendations
A number of methodological issues make it difficult to definitively determine the public health risks of drinking reclaimed water.
Despite these uncertainties, any utility considering the implementation of a potable reuse project should estimate the increased risk from microbial and chemical contaminants in reclaimed water relative to those from other available sources of water.
Microbiological Methods and Risk Assessment
There is a lack of information nationwide on the levels of viral and protozoan pathogens in all waters and the efficacy of both conventional water treatment and wastewater treatment for water reclamation in reducing these levels. The Information Collection Rule, promulgated in 1996 by the EPA, is a first step toward providing some of the exposure data needed for more effective risk assessments, but additional steps are needed to improve methods for assessing risks posed by microbial pathogens in water reuse projects.
Potable reuse projects should consider using some of the newer analytical methods, such as biomolecular methods, as well as new in-
dicator microorganisms, such as Clostridium perfringens and the F-specific coliphage virus, to screen drinking water sources derived from treated wastewaters. The microbial methods currently used for detecting bacterial, viral, and protozoan pathogens in water all have limitations when used to detect pathogens in reclaimed water. Bacterial techniques do not account for viable, noncultivable bacteria, and the new techniques for assessing the viability of protozoan cysts or oocysts will require evaluation. Standard cell culture methods employing tests of cytopathic effects have been limited to the detection of well-known enteroviruses and do not account for many other enteric viruses that may be found in wastewater. New analytical methods for rapid measurement of health-related microbial contaminants, such as the polymerase chain reaction, are being developed. However, treatment performance and water quality goals have not been developed for these methods. In addition, several new indicator microorganisms are available, including Clostridium perfringens and the F-specific coliphage virus, and should be considered as alternatives to using the coliform bacteria as indicators for water quality. In particular, F-specific coliphage can be used in seeded studies to provide useful data on unit process removals.
The EPA should include data on the concentrations of waterborne pathogens in source water in the new Drinking Water National Contaminant Occurrence Data Base and should develop better data on reductions of waterborne pathogens by various levels of treatment. The lack of monitoring data for evaluating exposure remains the greatest single barrier to the development of risk assessment for microbial pathogens. Microbial risk assessment requires better estimates of exposure, which should be based on monitoring data, to identify the concentration of microbial pathogens in raw wastewater, wastewater treated with various processes, ambient water, and drinking water treated with various processes.
State officials, water utilities, and water research scientists should document survival rates of relevant protozoa in natural environments. Indirect potable reuse projects may rely on dilution in the environment and reduction by natural processes (i.e., die-off in ambient waters and removal by soil infiltration systems) to remove pathogens of all kinds; however, while reductions of bacteria and viruses have been well documented, the information on protozoa survival in ambient waters remains inadequate. More research is needed to fill that gap.
Risk estimates should consider the effects on sensitive populations and the potential for secondary spread of infectious disease within a community. This precaution is necessary to prevent pathogens from infecting sensitive populations (the elderly or very young, or those with
suppressed immune systems) in whom mortality may be high and from whom diseases might spread to others.
The research community should conduct further studies to document the removal of pathogens of all types by membrane processes. Membrane systems, such as microfiltration, ultrafiltration, nanofiltration, and reverse osmosis, show the potential for nearly complete rejection of pathogens above certain size classes (in the case of the latter three processes, all size classes). More work is needed to demonstrate the suitability of these processes for potable reuse applications and to develop monitoring methods capable of continuously assessing process performance.
Chemical Risk Assessment
Analysis of reclaimed water is complicated by the fact that toxicological data on the wide variety of organic compounds in wastewater are much less complete than those for inorganic compounds. Thus, even if the organic material in the water could be analyzed completely, there would be no basis for assigning risks to most of the identifiable compounds present. This situation complicates the management of risks from chemical contaminants.
A conventional toxicological safety testing strategy developed in the food and drug industries uses both live animal (in vivo) and cell culture (in vitro) testing. While this approach has been used to develop risk assessments and regulations for recognized chemical contaminants in drinking water, there has been little experience in applying the strategy to determine health risks posed by the poorly characterized mixtures of organic chemicals in reclaimed water. The current state of knowledge in toxicology is too limited to make meaningful risk assessments based on in vitro data alone. So far, most toxicological studies of potable reuse have focused on bacterial and/or in vitro mammalian tests of genotoxicity of product waters rather than comprehensive testing on live animals. These bacterial or in vitro tests do not accurately evaluate the risks posed by the complex mixtures of contaminants in reclaimed wastewater.
Because of the uncertainty in the organic composition of reclaimed water, toxicological testing should be the primary component of chemical risk assessments of potable reuse systems. Attempting to ensure the safety of reclaimed water by analyzing only for known chemical contaminants, such as those regulated under the Safe Drinking Water Act, will not provide adequate protection of public health.
In waters where toxicological testing appears to be important for determining health risks, emphasis should be placed on live animal test systems capable of expressing a wide variety of toxicological ef-
fects. Chapter 5 presents a proposed system using fish in ambient waters.
Further, toxicological testing standards for reclaimed water should be supplemented by strict regulation of the processes for ''manufacturing" the water. Regulators should review the processes for manufacturing the reclaimed water (that is, the treatment systems and environmental storage employed) on a plant-by-plant basis.
Epidemiological Methods
Several methodological challenges complicate epidemiologic investigations of the health effects of potable water reuse. These challenges include (1) obtaining accurate measures of individual or group exposure to the waterborne agent of interest, (2) selecting the appropriate health outcomes to monitor, (3) accurately measuring health outcomes through either a surveillance system or individual health records, (4) estimating the fraction of disease cases due to waterborne exposure, (5) selecting an appropriate comparison group, and (6) recruiting a study population large enough to detect a true effect.
Given these challenges, the results of epidemiologic studies should be interpreted with caution and a recognition of the potential for systematic and random error and potential biases. The strongest observational study design for establishing a cause-and-effect relationship between exposure to waterborne disease agents and disease occurrences is a cohort study, which compares the disease rates over time among individuals who are exposed to reclaimed water to disease rates among individuals who use a different water source.
References
Asano, T., L. Y. C. Leong, M. G. Rigby, and R. H. Skaaji. 1992. Evaluation of the California wastewater reclamation criteria using enteric virus monitoring data. Water Science and Technology 26:1513-1524.
Ashby, J., and R. W. Tennant. 1988. Chemical structure, Salmonella mutagenicity and extent of carcinogenicity as indicators of genotoxic carcinogenesis among 222 chemicals tested in rodents by U.S. NCI/NTP. Mutation Research 204:17-115.
Atlas, R. M., G. Sayler, R. S. Burlage, and A. K. Bej. 1992. Molecular approaches for environmental monitoring of microorganisms. BioTechniques 12:706-717.
Bailey, N. T. J. 1975. The Mathematical Theory of Infectious Diseases and Its Applications. New York: Oxford University Press.
Bailey, N. T. J. 1986. Macro-modeling and prediction of epidemic spread at community level. Mathematical Modeling 7: 698-717.
Baron, R. C., F. D. Murphy, H. B. Greenberg, C. E. Davis, D. J. Bergman, G. W. Gary, J. M. Hughes, and L. B. Schonberher. 1982. Norwalk gastrointestinal illness: an outbreak associated with swimming in a recreational lake and secondary person to person transmission. American Journal of Epidemiology 115(2):163-172.
Bej, A K., M. H. Mahbubani, R. Miller, J. L. DiCesare, L. Haff, and R. M. Atlas. 1990. Multiplex PCR amplification and immobilized capture probes for detection of bacterial pathogens and indicators in water. Molecular and Cellular Probes 4:353-365.
Benenson, A.S. (ed.) 1995. Control of Communicable Diseases Manual, 16 ed. Washington, D.C.: American Public Health Association.
Cabelli, V. J.1997. Clostridium perfringens as a water quality indicator. Pp. 247-264 in Hoadley, A. W., and B. J Durka (eds.) Bacterial Indicators/Health Hazards Associated With Water. ASTM STP 635. Philadelphia: American Society for Testing and Materials.
Cabelli, V. J., A. P. Dufour, M. A. Levin, L. J. McCabe, and P. W. Haberman. 1979. Relationship of microbial indicators to health effects in marine bathing beaches. American Journal of Public Health 69(7):690-696.
Casanova, M., D. Bell, and H. Heck. 1997. Dichloromethane metabolism to formaldehyde (HCHO) and reaction of HCHO with nucleic acids in hepatocytes of rodents and humans with and without glutathione S-transferase T1 and M1 genes: adduct dosimetry and risk assessment. Submitted for publication.
CH2M Hill. 1993. Tampa Water Resource Recovery Project. Tampa, Flo.: CH2M Hill.
Clancy, J. L., W. D. Gollnitz, and Z. Tabib. 1994. Commercial labs: how accurate are they? Journal of the American Water Works Association 86:89-97.
Clausen, E. A.,B. L. Green, and W. Litsky. 1977. Fecal streptococci: indicator of pollution. Pp. 247-264 in Hoadley, A. W., and B. J. Dutka (eds.) Bacterial Indicators/Health Hazards Associated With Water. ASTM STP 635. Philadelphia: American Society for Testing and Materials.
Cohen, J., and H. I. Shuval. 1973. Water, Air, and Soil Pollution 2:85-95. New York, N. Y.: Springer-Verlag.
Coin, L. 1966. Modern microbiology and virological aspects of water pollution. Pp. 1-10 in Jaag, O. (ed.) Advances in Water Pollution Research. London: Pergamon Press.
Craun, G. F. 1993. Epidemiology Studies of Water Disinfectants and Disinfection Byproducts. Pp. 277-301 in Craun, G. F. (ed.) Safety of Water Disinfection: Balancing Chemical and Microbial Risks. Washington, DC: ILSI Press
Craun, G. F., R. L. Calderon, and F. J. Frost. 1996. An introduction to epidemiology. Journal of the American Water Works Association 88:54-65.
Davies-Colley, R. J., R. J. Bell, and A. M. Donnison. 1994. Sunlight inactivation of enterococci and fecal coliforms in sewage effluent diluted in seawater. Applied Environmental Microbiology 60:2049-2058.
Deng, M. Y., S. P. Day, and D. O. Cliver. 1994. Detection of hepatitis A virus in environmental samples by antigen-capture PCR. Applied and Environmental Microbiology 60:1927-1933.
Douglas, G. R., D. H. Blakely, and D. B. Clayson. 1988. IPCEMC Working Paper No. 5: Genotoxicity tests as predictors of carcinogens: An analysis. Mutation Res. 196: 83-93.
Dufour, A. P. 1984. Bacterial indicators of recreational water quality. Canadian Journal of Public Health 75:49-56.
Eisenberg, J., E. Seto, A. W. Olivieri, and R. C. Spear. 1996. Quantifying water pathogen risk in an epidemiological framework. Risk Analysis 16(4):549-563.
Fujioka, R. S., and L. K. Shizumura. 1985. Clostridium perfringens, a reliable indicator of stream water quality. Journal of the Water Pollution Control Federation 57:986-992.
Fung, D. Y. C. 1994. Rapid methods and automation in food microbiology: a review. Food Reviews International 10(3): 357-375.
Garcia, L. S., T. C. Brewer, and A. Bruckner. 1987. Fluorescence detection of Cryptosporidium oocysts in human fecal specimens by using monoclonal antibodies. Journal of Clinical Microbiology 25:119-121.
Gerba, C. P., and J. B. Rose. 1990. Viruses in source and drinking water. Pp. 380-396 in McFeters, G. A. (ed.) Drinking Water Microbiology. New York: Springer-Verlag.
Gerba, C. P., J. B. Rose, and C. N. Haas. 1996. Sensitive populations:who is at greatest risk? International Journal of Food Microbiology 30(1-2):113-123.
Greenberg, R. S. 1993. Medical Epidemiology. Norwalk, Conn.: Appleton and Lange.
Griffin, P. M., and R. V. Tauxe. 1991. The epidemiology of infections caused by Escherichia coli 0157:h7, other enterohemorrhagic E. coli and the associated hemolytic uremic syndrome. Epidemiologic Reviews 13: 60-98.
Haas, C. N. 1983. Estimation of risk due to low doses of microorganisms: A comparison of alternative methodologies. American Journal of Epidemiology 118:573-582.
Haas, C. N., J. B. Rose, C. P. Gerba, and S. Regli. 1993. Risk assessment of viruses in drinking water. Risk Analysis 13:545-552.
Haas, C. N., C. Crockett, J. B. Rose, C. Gerba, and A. Fazil. 1996. Infectivity of Cryptosporidium parvum oocysts. Journal of the American Water Works Association 88(9):131-136.
Havelaar, A. H., M. Van Olphen, and Y. C. Drost. 1993. F-specific RNA bacteriophages are adequate model organisms for enteric viruses in fresh water. Applied and Environmental Microbiology 59:2956-2962.
Hurst, C. J., K. A. McClellan, and W. H. Benton. 1988. Comparison of cytopathogenicity, immunofluorescence and in situ DNA hybridization as methods for the detection of adenoviruses. Water Research 22:1547-1552.
Hurst, C. J., W. H. Benton, and R. E. Stetler. 1989. Detecting viruses in water. Journal of the American Water Works Association 9:71-80.
ILSI. 1996. Disinfection by-products in drinking water: critical issues in health effects research. Pp. 110-120 in Workshop Report. Chapel Hill, N.C.: October 23-25, 1995.
Jakubowski, W., S. Boutros, W. Faber, R. Fayer, W. Ghiorse, M. LeChevallier, J. Rose, S. Schaub, A. Singh, and M. Stewart. 1996. Environmental methods for Cryptosporiodium. Journal of the American Water Works Association 88(9):107-121.
Johnson, D. W., N. J. Pieniazek, D. W. Griffin, L. Misener, and J. B. Rose. 1995. Development of a PCR protocol for sensitive detection of Cryptosporidium in water samples. Applied and Environmental Microbiology 61(11):3849-3855.
Kopecka, H., S. Dubrou, J. Prevot, J. Marechal, and J. M. Lopez-Pila. 1993. Detection of naturally occurring enteroviruses in waters by reverse transcription, polymerase chain reaction, and hybridization. Applied and Environmental Microbiology 59:1213-1219.
Lauer, W. C., F. J. Johns, G. W. Wolfe, B. A. Meyers, L. W. Condie, and J. F. Borzelleca. 1990. Comprehensive health effects testing program for Denver's potable reuse demonstration project. J. Toxicol. Environ. Health 30:305-321.
LeChevallier, M. W., and W. D. Norton. 1993. Treatment to address source water concerns: protozoa. Pp. 145-164 in Craun, G. F. (ed.) Safety of Water Disinfection: Balancing Chemical and Microbial Risks. Washington, D.C.: ILSI Press.
LeChevallier, M. W., and T. M. Trok. 1990. Comparison of the zinc sulfate and immunofluorescence techniques for detecting Giardia and Cryptosporidium. Journal of the American Water Works Association 82:75-82.
LeChevallier, M. W., W. D. Norton, and R. G. Lee. 1991a. Giardiaand Cryptosporidium in filtered drinking water supplies . Applied and Environmental Microbiology 57:2617-2621.
LeChevallier, M. W., W. D. Norton, and R. G. Lee. 1991b. Occurrence of Cryptosporidium and Giardia spp in surface water supplies. Applied and Environmental Microbiology 57:2610-2616.
Linquist, H. A. D. 1997. Probes for specific detection of Cryptosporidium parvum. Wat. Res. 31(10):2668-2671.
MacKenzie, W. R., N. J. Hoxie, M. E. Proctor, M. S. Gradus, K. A. Blair, D. E. Peterson, J. J. Kazmierczak, D. G. Addiss, K. R. Fox, and J. B. Rose. 1994. A massive outbreak in Milwaukee of Cryptosporidium infection transmitted through the public water supply. New England Journal of Medicine 331:161-167.
MacKenzie, W. R., W. L. Schell, K. A. Blair, D. G. Addiss, D. E. Peterson, N. J. Hoxie, J. J. Kazmierczak, and J. P. Davis. 1995. Massive outbreak of waterborne Cryptosporidium infection in Milwaukee, Wisconsin: recurrence of illness and risk of secondary transmission. Clinical Infectious Diseases 21:57-62.
Mahbubani, M. H., A. K. Bej, M. Perlin, F. W. Schaefer, W. Jakubowski, and R. M. Atlas. 1991. Detection of Giardia cysts by using the polymerase chain reaction and distinguishing live from dead cysts. Applied and Environmental Microbiology 57:3456-3461.
Millard, P.S., K. F. Gensheimer, D. G. Addiss, D. M. Sosin, G. A. Beckett, A. Houck-Jankoski, and A. Hudson. 1994. An outbreak of cryptosporidiosis from fresh-pressed apple cider. Journal of the American Medical Association 272(20): 1592-1596.
Morens, D. M., R. M. Zweighaft, T. M. Vernon, G. W. Gary, J. J. Eslein, B. T. Wood, R. C. Holman, and R. Dolin. 1979. A waterborne outbreak of gastroenteritis with secondary person to person spread. Lancet May 5: 964-966.
National Institutes of Health (NIH). 1993. A chemical mixture of 25 groundwater contaminants. National Toxicology Program. NIH Report 93-3384. Research Triangle Park, NC: NIH.
National Research Council (NRC) 1977-1989. Drinking Water and Health, vols. 1-9. Washington, D.C.: National Academy Press.
National Research Council (NRC). 1982. Quality Criteria for Water Reuse. Washington, D.C.: National Academy Press.
National Research Council (NRC). 1983. Risk Assessment in the Federal Government: Managing the Process. Washington, D.C.: National Academy Press.
National Research Council (NRC). 1989. Drinking Water and Health: Selected Issues in Risk Assessment, Vol. 9. Washington, D.C.: National Academy Press.
National Research Council (NRC). 1994. Science and Judgment in Risk Assessment. Washington, D.C.: National Academy Press.
National Research Council (NRC). 1996. Use of Reclaimed Water and Sludge in Food Crop Production. Washington, D.C.: National Academy Press.
New York City's Advisory Panel on Waterborne Disease Assessment. 1994. Report of New York City's Advisory Panel on Waterborne Disease Assessment. October 7. The New York City Department of Environmental Protection.
Ongerth, J. E., and H. H. Stibbs. 1987. Identification of Cryptosporidium oocysts in river water. Applied and Environmental Microbiology 53: 672-679.
Parodi, S., M. Taning, P. Russo, M. Pala, D. Vecchio, G. Fassina, and L. Santi. 1983. Quantitative predictivity of the transformation in vitro assay compared with the Ames test. J. Toxicol. Environ. Health 12:483-510.
Paul, J. R., and J. D. Trask. 1947. The virus of poliomyelitis in stools and sewage. J. Am. Med. Assoc. 116:493-497.
Paul, J. H., J. B. Rose, J. Brown, E. A. Shinn, S. Miller, and S. Farrah. 1995. Viral tracer studies indicate contamination of marine waters by sewage disposal practices in Key Largo, Florida. Applied Environmental Microbiology 61:2230-2234.
Payment, P., and E. Franco. 1993. Clostridium perfringens and somatic coliphages as indicators of the efficiency of drinking water treatment for viruses and protozoan cysts. Applied and Environmental Microbiology 59:2418-2424.
Payment, P., and M. Trudel. 1985. Detection and health risk associated with low virus concentration in drinking water. Water Science and Technology 17:97-103.
Payment, P., M. Trudel, and R. Plante. 1985. Elimination of viruses and indicator bacteria at each step of treatment during preparation of drinking water at seven water treatment plants. Applied and Environmental Microbiology 49:1418-1428.
Payment, P., L. Richardson, J. Siemiatycki, R. Dewar, M. Edwardes, and E. Franco. 1991. A randomized trial to evaluate the risk of gastrointestinal disease due to consumption of drinking water meeting current microbiological standards. American Journal of Public Health 81(6):703-708.
Pearce, N., S de Sanjose, P. Boffetta, M. Kogevinas, R. Saracci, and D. Savitz. 1995. Limitations of biomarkers of exposure in cancer epidemiology. Epidemiology 6:190-194.
Pickering, L. K., D. G. Evans, H. L. DuPont, J. J. Vollet, III, and D. J. Evans, Jr. 1981. Diarrhea caused by shigella, rotavirus and giardia in day care centers: prospective study. Journal of Pediatrics 99(1): 51-56.
Powelson, D. K., and C. P. Gerba. 1995. Fate and transport of microorganisms in the vadose zone. Pp. 123-135 in Wilson, L. G., L. G. Everett, and S. J. Cullen (eds.) Handbook of Vadose Zone Characterization and Monitoring. Boca Raton, Flo.: Lewis Publishers.
Regli, S., J. B. Rose, C. N. Haas, and C. P. Gerba. 1991. Modeling the risk of Giardia and viruses in drinking water. Journal of the American Water Works Association 83:7684.
Rose, J. B. 1997. Environmental ecology of Cryptosporidium and public health implications. Ann. Rev. Pub. Health 18: 135-161.
Rose, J. B., D. Kayed, M. S. Madore, C. P. Gerba, M. J. Arrowood, and C. R. Sterling. 1988a. Methods for the recovery of Giardia and Cryptosporidium from environmental waters and their comparative occurrence. In Wallis, P., and B. Hammond (eds.) Advances in Giardia Research. Calgary: University of Calgary Press.
Rose, J. B., H. Darbin, and C. P. Gerba. 1988b. Correlations of the protozoa, Cryptosporidium and Giardia with water quality variables in a watershed. Water Science and Technology 20:271-276.
Rose, J. B., L. K. Landeen, K. R. Riley, and C. P. Gerba. 1989. Evaluation of immunofluorescence techniques for detection of Cryptosporidium oocysts and Giardia cysts from environmental samples. Applied and Environmental Microbiology 55:3189-3195.
Rose, J. B., C. N. Haas, and S. Regli. 1991a. Risk assessment and control of waterborne giardiasis. American Journal of Public Health 81:709-713.
Rose, J. B., C. P. Gerba, and W. Jakubowski. 1991b. Survey of potable water supplies for cryptosporidium and giardia. Environmental Science and Technology 25:1393-1400.
Rose, J. B., L. J. Dickson, S. R. Farrah, and R. P. Carnahan. 1996. Removal of pathogenic and indicator microorganisms by a full scale water reclamation facility. Water Research 30(11):2785-2797.
Savitz, D., and C. L. Moe. 1997. Water: chlorinated hydrocarbons and infectious agents. Pp. 89-118 in Steenland, N. K., and D. A. Savitz (eds.) Topics in Environmental Epidemiology. New York: Oxford University Press.
Schulte, P. A. 1993. A conceptual and historical framework for molecular epidemiology. Pp 3-44 in Schulte, P., and F. R. Perera (eds.) Molecular Epidemiology: Principles and Practices. New York: Academic Press.
Singh, A., and G. A. McFeters. 1990. Injury of Enteropathogenic bacteria in drinking water. Pp. 368-379 in McFeters, G. A. (ed.) Drinking Water Microbiology. New York: Springer-Verlag.
Sinton, L. W., R. J. Davies-Colley, and R. G. Bell. 1994. Inactivation of enterococci and fecal coliforms from sewage and meatworks effluents in seawater chambers. Applied Environmental Microbiology 60:2040-2048.
Slifko, T. E., D. E. Friedman, J. B. Rose, and W. Jakubowski. 1997. An in vitro method for detecting infectious Cryptosporidium oocysts with cell culture. Applied and Environmental Microbiology 63(9): 3669-3675.
Sloss, E. M., S. A. Geschwind, D. F. McCaffrey, and B. R. Ritz. 1996. Groundwater Recharge with Reclaimed Water: An Epidemiologic Assessment in Los Angeles County, 1987-1991. Santa Monica, Calif.: RAND.
Smith, J. L., S. A. Palumbo, and I. Walls. 1993. Relationship between foodborne bacterial pathogens and the reactive athritides. Journal of Food Safety 13:209-236.
Smith, G., L. Stanley, E. Sim, R. C. Strange, and C. R. Wolf. 1995. Metabolic polymorphisms and cancer susceptibility. Cancer Survey 25:27-65.
Spika, J. S., J. E. Parsons, D. Nordenberg, J. D. Wells, R. A. Gunn, and P. A. Blake. 1986. Hemolytic uremic syndrome and diarrhea associated with Escherichia coli 0157:h7 in a day care center. Journal of Pediatrics 109: 287-291.
Stetler, R. E., R. L. Ward, and S. C. Waltrip. 1984. Enteric virus and indicator bacteria levels in a water treatment system modified to reduce trihalomethane production. Applied and Environmental Microbiology 47:319-324.
Stibbs, H. H., E. T. Riley, J. Stockard, J. Riggs, P. M. Wallis, and J. Issac-Renton. 1988. Immunofluorescence differentiation between various animal and human source Giardia cysts using monoclonal antibodies. Pp. 159-163 in Wallis, P., and B. Hammond (eds.) Advances in Giardia Research. Calgary: University of Calgary Press.
Sutter, T. E. 1995. Molecular and cellular approaches to extrapolation for risk assessment. Environ. Health Persp. 103:386-389.
Thier, R., J. B. Taylor, S. E. Pemble, W. G. Humphreys, M. Persmark, B. Ketterer, and F. P. Guengerich. 1993. Expression of mammalian glutathione S-transferase 5-5 in Salmonella typhimurium TA1535 leads to base-pair mutations upon exposure to dihalomethanes. Proc. Natl. Acad. Sci. 90:8576-80.
Tsai, Y. L., and B. H. Olson. 1991. Rapid method for direct extraction of DNA from soil and sediments. Applied and Environmental Microbiology 57 (4):1070-1074.
Tsai, Y. L., and B. H. Olson. 1992a. Detection of low numbers of bacterial cells in soils and sediments by polymerase chain reaction. Applied and Environmental Microbiology 58:754-757.
Tsai, Y. L., and B. H. Olson. 1992b. Rapid method for separation of bacterial DNA from humic substances in sediments for polymerase chain reaction. Applied and Environmental Microbiology 58:2292-2295.
U.S. Environmental Protection Agency (EPA). 1986. Ambient Water Quality Criteria for Bacteria. EPA 44015-84-002. Office of Regulations and Standards. Washington, D.C.: U.S. EPA.
U.S. Environmental Protection Agency (EPA). 1989a. Surface water treatment rule guidance manual for compliance with the filtration and disinfection requirements for public water systems using surface water sources. Federal Register 54(124). June 29.
U.S. Environmental Protection Agency (EPA). 1989b. Surface water treatment rule guidance manual for compliance with the filtration and disinfection requirements for public water systems using surface water sources. Federal Register 54(124) June 29.
U.S. Environmental Protection Agency (EPA). 1989c. National primary drinking water regulation; filtration and disinfection; turbidity; Giardia lamlia; viruses, Legionella and heterotophic bacteria. Federal Register 54:27486-27541.
U.S. Environmental Protection Agency (EPA). 1992. Guidelines for Water Reuse. EPA 625/ R-92/004. Washington, D.C.: U.S. EPA.
U.S. Environmental Protection Agency (EPA). 1996. Proposed cancer risk assessment guidelines. EPA/600/P-92/003C. Washington, D.C.














































