Important Points Highlighted by Individual Speakers
• Genomic information has great potential to identify new path-ways involved in complex diseases, suggest new therapeutic targets, evaluate adverse drug effects, and identify populations for which a drug is most effective or has the least deleterious effects.
• Pharmaceutical and biotechnology companies have integrated-based genomics-based strategies for drug discovery, but this has largely not been translated into late-stage development.
• The cost of therapeutic development has increased significantly over the past few decades while the success rate has remained unchanged, and many drug failures often occur after large investments have been made.
• While targeted therapeutics may decrease market size, overall market share may increase, leading to a significant potential advantage for developing stratified medicines.
• Commercial and marketing organizations may need to be aligned with research and development in order to develop a successful commercial model for targeted therapeutics.
The advent of the genomic era generated great expectations for drug discovery and development, said Geoffrey Ginsburg of Duke University. Genomic information was expected to provide insights into the underlying biological mechanisms of disease and to highlight biological targets and pathways that would be amenable to new drug discovery. It indicated an approaching ability to stratify populations based on genomic-based biomarkers, leading to better clinical development programs. Genomic data would reveal how individuals might respond to, be resistant to, or have adverse effects from a drug, creating the potential for personalized medicines. As a result of these and other changes, genomic data would increase the efficiency of drug discovery and development, increase the success rate of new drugs, enhance safety, and decrease costs.
The genomic era has made major strides toward delivering on these promises, Ginsburg said. Several genomics-enabled products have been approved in recent years or are in development for use, including three that are described in Chapter 3 of this report: crizotinib for the treatment of non-small-cell lung cancer, pomaglumetad methionil for schizophrenia, and ivacaftor for cystic fibrosis. In addition, academic–industry partnerships have formed to leverage a deep understanding of disease biology from the academic realm and to meld that to product development and commercialization in industry. Precompetitive collaborations, such as the European Innovative Medicines Initiative and programs sponsored by the National Center for Advancing Translational Sciences (NCATS) at the National Institutes of Health (NIH), have sought to lay the groundwork for new therapeutics.
Nicholas Davies from PricewaterhouseCoopers (PwC) elaborated on the potential and current use of genomic-based drug discovery and development during his presentation in the workshop’s initial session. The efficiency and quality of research inputs have undergone huge improvements. The cost of DNA sequencing has dropped by many orders of magnitude and continues to drop. The ability to find targets, screen compounds, and generate chemical libraries is immense. The development of companion diagnostics has made it possible to target patient subpopulations that would be expected to benefit from a specific treatment. As Mark Trusheim from the Sloan School of Management at the Massachusetts Institute of Technology added, in this way patients and providers have more and better treatment options, regulators gain a better sense of risk-benefit comparisons, drug and diagnostic innovators generate more products and profits, and
payers spend less on ineffective therapies. “We see opportunities not just for developers and patients, but for everyone in the cycle. … It has to work for everyone or it is not going to work at all as a system.”
Both Ginsburg and Davies said that a genomic-based approach continues to have tremendous potential. For example, a recent analysis of genome-wide association studies (GWAS) found that such studies can reveal new pathways involved in complex diseases and suggest potential therapeutic options that had not previously been considered for those indications (Collins, 2011). This analysis also suggested that the off-target or adverse effects of those drugs could be monitored through the analysis of genes discovered through these unbiased genome-wide approaches.
Garret FitzGerald of the University of Pennsylvania added that it has already been demonstrated that genetic information can be used to evaluate adverse drug effects. Studies designed specifically to determine whether particular gene variants can be used to identify individuals at particular risk have been successful for both lumiracoxib and abacavir and required only very small numbers of study participants to do so.
According to recent data from the U.S. Food and Drug Administration (FDA), more than 110 marketed drugs have pharmacogenetic biomarkers on the label (see Table 2-1),1 and the need for further drugs developed through a genomic-based approach remains strong. As Trusheim observed, many major drugs, including hypertension drugs, heart failure drugs, antidepressants, cholesterol drugs, and asthma drugs, are ineffective for large portions of the population (Spear et al., 2001). Furthermore, ineffective therapies cause substantial harm. Medication-related health problems account for an estimated 3 to 7 percent of hospital admissions (Pirmohamed et al., 2004), and 15 percent of patients experience an adverse drug reaction during hospital stays. An important consequence of these adverse reactions is heightened patient noncompliance.
Oncology has made the most progress in developing personalized medicine (defined in Box 2-1), Davies said, but genomic-based research is also starting to make progress on diseases of the cardiovascular system, central nervous system, and immune system. Metabolic, respiratory, and viral diseases also are starting to yield to this approach, though progress has been slower than expected.
Pharmaceutical companies and biotechnology companies are striving to modernize their drug discovery and development processes. Davies pointed to data from the Tufts Center for the Study of Drug Development (Tufts, 2010) showing that 100 percent of surveyed companies are using a discovery strategy that involves a genetic or genomic approach. Thirty percent
![]()
1 For an up-to-date listing of these drugs, see http://www.fda.gov/Drugs/ScienceResearch/ResearchAreas/Pharmacogenetics/ucm083378.htm.
TABLE 2-1 Pharmacogenomic Biomarkers in Drug Labels
|
|
||
| Drug | Therapeutic Area | Biomarker |
|
|
||
|
Abacavir |
Antivirals |
HLA-B*5701 |
|
Aripiprazole |
Psychiatry |
CYP2D6 |
|
Arsenic Trioxide |
Oncology |
PML/RARα |
|
Atomoxetine |
Psychiatry |
CYP2D6 |
|
Atorvastatin |
Metabolic and Endocrinology |
LDL receptor |
|
Azathioprine |
Rheumatology |
TPMT |
|
Boceprevir |
Antivirals |
IL28B |
|
Brentuximab Vedotin |
Oncology |
CD30 |
|
Busulfan |
Oncology |
Ph Chromosome |
|
Capecitabine |
Oncology |
DPD |
|
Carbamazepine |
Neurology |
HLA-B*1502 |
|
Carisoprodol |
Musculoskeletal |
CYP2C19 |
|
Carvedilol |
Cardiovascular |
CYP2D6 |
|
Celecoxib |
Analgesics |
CYP2C9 |
|
Cetuximab (1) |
Oncology |
EGFR |
|
Cetuximab (2) |
Oncology |
KRAS |
|
Cevimeline |
Dermatology and Dental |
CYP2D6 |
|
Chlordiazepoxide and Amitriptyline |
Psychiatry |
CYP2D6 |
|
Chloroquine |
Anti-Infectives |
G6PD |
|
Cisplatin |
Oncology |
TPMT |
|
Citalopram (1) |
Psychiatry |
CYP2C19 |
|
Citalopram (2) |
Psychiatry |
CYP2D6 |
|
Clobazam |
Neurology |
CYP2C19 |
|
Clomiphene |
Reproductive and Urologic |
Rh genotype |
|
Clomipramine |
Psychiatry |
CYP2D6 |
|
Clopidogrel |
Cardiovascular |
CYP2C19 |
|
Clozapine |
Psychiatry |
CYP2D6 |
|
Codeine |
Analgesics |
CYP2D6 |
|
Crizotinib |
Oncology |
ALK |
|
Dapsone |
Dermatology and Dental |
G6PD |
|
Dasatinib |
Oncology |
Ph Chromosome |
|
Denileukin Diftitox |
Oncology |
CD25 |
|
Desipramine |
Psychiatry |
CYP2D6 |
|
Dexlansoprazole (1) |
Gastroenterology |
CYP2C19 |
|
Dexlansoprazole (2) |
Gastroenterology |
CYP1A2 |
|
Dextromethorphan and Quinidine |
Neurology |
CYP2D6 |
|
Diazepam |
Psychiatry |
CYP2C19 |
|
Doxepin |
Psychiatry |
CYP2D6 |
|
Drospirenone and Ethinyl Estradiol |
Reproductive |
CYP2C19 |
|
Erlotinib |
Oncology |
EGFR |
|
Esomeprazole |
Gastroenterology |
CYP2C19 |
|
Everolimus |
Oncology |
Her2/neu |
|
Exemestane |
Oncology |
ER &/PgR receptor |
|
|
||
|
Drug |
Therapeutic Area |
Biomarker |
|
|
||
|
Fluorouracil |
Dermatology and Dental |
DPD |
|
Fluoxetine |
Psychiatry |
CYP2D6 |
|
Fluoxetine and Olanzapine |
Psychiatry |
CYP2D6 |
|
Flurbiprofen |
Rheumatology |
CYP2C9 |
|
Fluvoxamine |
Psychiatry |
CYP2D6 |
|
Fulvestrant |
Oncology |
ER receptor |
|
Galantamine |
Neurology |
CYP2D6 |
|
Gefitinib |
Oncology |
EGFR |
|
Iloperidone |
Psychiatry |
CYP2D6 |
|
Imatinib (1) |
Oncology |
C-Kit |
|
Imatinib (2) |
Oncology |
Ph Chromosome |
|
Imatinib (3) |
Oncology |
PDGFR |
|
Imatinib (4) |
Oncology |
FIP1L1-PDGFRα |
|
Imipramine |
Psychiatry |
CYP2D6 |
|
Indacaterol |
Pulmonary |
UGT1A1 |
|
Irinotecan |
Oncology |
UGT1A1 |
|
Isosorbide and Hydralazine |
Cardiovascular |
NAT1; NAT2 |
|
Ivacaftor |
Pulmonary |
CFTR (G551D) |
|
Lapatinib |
Oncology |
Her2/neu |
|
Lenalidomide |
Hematology |
Chromosome 5q |
|
Letrozole |
Oncology |
ER &/PgR receptor |
|
Maraviroc |
Antivirals |
CCR5 |
|
Mercaptopurine |
Oncology |
TPMT |
|
Metoprolol |
Cardiovascular |
CYP2D6 |
|
Modafinil |
Psychiatry |
CYP2D6 |
|
Nefazodone |
Psychiatry |
CYP2D6 |
|
Nilotinib (1) |
Oncology |
Ph Chromosome |
|
Nilotinib (2) |
Oncology |
UGT1A1 |
|
Nortriptyline |
Psychiatry |
CYP2D6 |
|
Omeprazole |
Gastroenterology |
CYP2C19 |
|
Panitumumab (1) |
Oncology |
EGFR |
|
Panitumumab (2) |
Oncology |
KRAS |
|
Pantoprazole |
Gastroenterology |
CYP2C19 |
|
Paroxetine |
Psychiatry |
CYP2D6 |
|
Peginterferon alfa-2b |
Antivirals |
IL28B |
|
Perphenazine |
Psychiatry |
CYP2D6 |
|
Pertuzumab |
Oncology |
Her2/neu |
|
Phenytoin |
Neurology |
HLA-B*1502 |
|
Pimozide |
Psychiatry |
CYP2D6 |
|
Prasugrel |
Cardiovascular |
CYP2C19 |
|
Pravastatin |
Metabolic and Endocrinology |
ApoE2 |
|
Propafenone |
Cardiovascular |
CYP2D6 |
|
Propranolol |
Cardiovascular |
CYP2D6 |
|
Protriptyline |
Psychiatry |
CYP2D6 |
|
Quinidine |
Antiarrhythmics |
CYP2D6 |
|
Rabeprazole |
Gastroenterology |
CYP2C19 |
|
|
||
|
Drug |
Therapeutic Area |
Biomarker |
|
|
||
|
Rasburicase |
Oncology |
G6PD |
|
Rifampin, Isoniazid, and Pyrazinamide |
Anti-Infectives |
NAT1; NAT2 |
|
Risperidone |
Psychiatry |
CYP2D6 |
|
Sodium Phenylacetate and Sodium Benzoate |
Gastroenterology |
UCD (NAGS; CPS; ASS; OTC; ASL; ARG) |
|
Sodium Phenylbutyrate |
Gastroenterology |
UCD (NAGS; CPS; ASS; OTC; ASL; ARG) |
|
Tamoxifen |
Oncology |
ER receptor |
|
Telaprevir |
Antivirals |
IL28B |
|
Terbinafine |
Antifungals |
CYP2D6 |
|
Tetrabenazine |
Neurology |
CYP2D6 |
|
Thioguanine |
Oncology |
TPMT |
|
Thioridazine |
Psychiatry |
CYP2D6 |
|
Ticagrelor |
Cardiovascular |
CYP2C19 |
|
Tolterodine |
Reproductive and Urologic |
CYP2D6 |
|
Tositumomab |
Oncology |
CD20 antigen |
|
Tramadol and Acetaminophen |
Analgesics |
CYP2D6 |
|
Trastuzumab |
Oncology |
Her2/neu |
|
Tretinoin |
Dermatology and Dental |
PML/RARα |
|
Trimipramine |
Psychiatry |
CYP2D6 |
|
Valproic Acid |
Psychiatry |
UCD (NAGS; CPS; ASS; OTC; ASL; ARG) |
|
Vemurafenib |
Oncology |
BRAF |
|
Venlafaxine |
Psychiatry |
CYP2D6 |
|
Voriconazole |
Antifungals |
CYP2C19 |
|
Warfarin (1) |
Hematology |
CYP2C9 |
|
Warfarin (2) |
Hematology |
VKORC1 |
|
|
||
SOURCE: U.S. Food and Drug Administration.
“Personalized medicine” or “stratified medicine,” as used by speakers in the workshop, refers to the use of an individual’s characteristics, including genetic information, to guide medical decisions regarding prevention, diagnosis, and treatment of disease. This tailoring of medical treatments is based on the ability to classify individuals into subpopulations so that they can benefit from the most efficacious treatments or interventions or be spared from expense or deleterious side effects.
require that all their compounds have an associated biomarker before going into clinical development. More than 80 percent of companies have established strategic partnerships related to personalized medicine, and half have collected DNA samples from clinical trial participants. In addition, companies that have developed genomic and diagnostic technologies have recently been acquired by other companies, suggesting that these approaches continue to be viewed favorably. However, Davies said, in large part these methods are not being employed in late-stage development due to a reluctance on the part of pharmaceutical companies to enable genomic-or genetic-based trials.
One concern about targeted drugs, Trusheim said, has been that they will have smaller markets and therefore attract less investment. But higher efficacy for targeted groups can in fact yield more market share and help minimize the overall reduction in market size (Figure 2-1; Trusheim et al., 2007). Underserved patients may enter the market and look for treatment
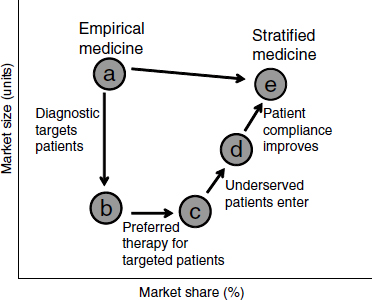
NOTE: As defined by Trusheim et al. (2007), an empirical medicine, as opposed to a stratified medicine, is not developed based upon the characteristics of an individual or a subpopulation of individuals. These medicines are based on overall population response and may work for a large or a small amount of individuals without using (either because it is not necessary or one is not available) a methodology to identify which groups may respond.
SOURCE: Trusheim et al., 2007.
if they are more confident that a treatment will work for them. In addition, providers may be more confident to prescribe a drug, especially since possible side effects are outweighed by the benefits. If biomarkers can separate those who will respond from those who will not, a drug will perform much better in the response group, potentially leading to quicker adoption, better patient compliance, more market share, and a higher price premium. This can produce a “niche buster” where the clinical performance of the drug and diagnostic drives commercial performance. For example, a study of the use of trastuzumab and panitumumab in cancer and bapineuzumab in Alzheimer’s disease showed a substantial potential economic advantage to using stratified-medicine strategies (Trusheim et al., 2011).
A final consideration in the adoption of personalized medicine, Davies said, is that cost-effective and outcomes-driven therapy will be critical in the future as health care changes. Care will become more preventive, and medicine in general will be more patient-centric. Cost control and value in outcomes will be increasingly important focuses. New therapies may need to be cost neutral, in that they make up for the additional expense of the therapy through reduced costs elsewhere, whether hospital readmissions, surgery, or some other form of care.
Genomic-based approaches are an area of promise in an otherwise troubled industry. The success rate for new drugs in the pharmaceutical industry—with success defined as the ability to identify a compound that will be approved and be commercially successful—has remained more or less constant over the last few decades, with occasional upticks, Ginsburg noted (Mullard, 2012). On average, fewer than 1 in 10 compounds entering preclinical testing will be successful. Furthermore, as Davies observed, failures often occur after large investments have been made. In 2010, 45 separate drugs failed in Phase III clinical trials, with the average cost for a Phase III trial being about $100 million. Meanwhile, patents are expiring on profitable drugs, which is further reducing resources. The costs of failures add to development expense and decrease the willingness to invest in the process.
Because of declining productivity, more resources have been needed to produce a constant level of new drugs. According to an analysis in Nature Reviews Drug Discovery, productivity in the pharmaceutical industry, measured in terms of output per billion dollars spent, has been decreasing logarithmically (Figure 2-2). This declining productivity has become known as “Eroom’s law.” “Eroom” is “Moore” spelled backward, and the name is meant to imply a backward version of “Moore’s law,” the observation made by Intel co-founder Gordon Moore in 1965 that the number of
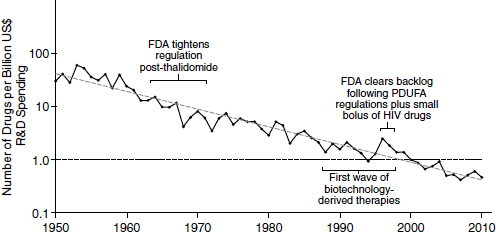
NOTE: FDA, U.S. Food and Drug Administration; HIV, human immunodeficiency virus; PDUFA, Prescription Drug User Fee Act; R&D, research and development.
SOURCE: Scannell et al., 2012.
components in integrated circuits was doubling approximately every year. In the pharmaceutical industry, the output per billion dollars spent has consistently decreased by half every 9 years since 1952.
To remain in the pharmaceutical business, companies and investors need to make money. But the return on capital investment is diminishing to the point that the existing financial model is no longer sustainable, Davies said. The average return on capital after 5 years’ sales is currently about $75 million per billion dollars invested, which is clearly not sustainable. A recent report in Forbes magazine estimated that some companies are spending upward of $12 billion per launched product (Herper, 2012). As FitzGerald noted, “catastrophe rather than opportunity usually drives radical change … and this model is about to change.”
According to Davies, the pharmaceutical industry invested an estimated $125 billion in research and development across the industry (Hewitt et al., 2011). An estimated 5 percent of this amount was spent specifically on genetic and genomic research, or about $6 billion including partnerships, acquisitions, and internal research. Companies have slightly different levels and strategies of investment, with some investing more heavily in internal research and some more heavily in external research. As discussed later in this summary and in a prior Roundtable on Translating Genomic-Based Research for Health workshop (IOM, 2011), academic partnerships have become popular, though ways of esti-
mating the value generated by these partnerships remain rudimentary. “Pharma[ceutical companies] and academia need to understand how to work together more effectively and demonstrate that they generate value from [partnering],” Davies said.
CHALLENGES FOR GENOMIC-BASED APPROACHES
Despite its promise, a genomic-based approach to drug discovery and development is surrounded by great uncertainties, as noted by each of the speakers in the workshop’s opening session. As FitzGerald pointed out, genomic testing must be shown to influence clinical outcomes to guarantee reimbursement. Adoption will also require substantial physician and patient education, a financial incentive for test development, and patent protection. Davies observed that oncology has been the poster child for a molecular approach to target discovery, diagnosis in the clinic, development of companion diagnostics, and treatment. However, these therapies tend to be expensive, making their value in general medicine uncertain. Furthermore, outside oncology, the value of targeted therapies for the most part remains to be determined. In addition, regulatory constraints are getting tighter, which is an issue for thinking about innovative approaches to bringing medicines to market with companion diagnostics or a targeted approach.
In general, Davies continued, the commercial model for the development of personalized medicines remains immature, with the commercial and marketing organizations within industry retaining a preference to go to market with a more general molecule than with a targeted therapeutic. An analysis by PwC estimates that the companion diagnostic market will reach $42 billion by 2015.2 “There is a huge market for companion diagnostics,” Davies said, “but they are culturally and from a time perspective [off]-kilter with the development cycle and culture of the research and development industry.”
Trusheim added that there are countervailing forces at play. Developing both a drug and a diagnostic can take longer, especially given the need to recruit targeted patient pools and synchronize development of the diagnostic; the resulting market may be smaller than for a more general drug; and developers face an increased risk of failure since the drug approval is dependent upon simultaneous approval of the diagnostic. Further complicating the matter, regulatory requirements differ because therapeutics and diagnostics generally fall under different legislative authorities. In addition, product exclusivity concerns raise profitability questions among companies.
![]()
2 For more information, see http://www.pwc.com/gx/en/pharma-life-sciences/pharmaceutical-industry-thought-leadership/pharma-life-sciences-mergers-acquisitions-diagnostics-2011.jhtml.
Patients are prone to confusion regarding the value of genetic and genomic technologies, and providers tend to be untrained in these areas, which slows adoption. Drug reimbursement has been slow, reimbursements for diagnostics remain focused on costs rather than value, and payers do not invest in research and development, even though they benefit from stratification. Many payers and health care providers remain unconvinced that many such therapies improve people’s health or are cost-effective.
In addition to its successes, Trusheim said, genomic-based drug development offers cases where these challenges have so far prevailed. For example, no candidate marker for response to bevacizumab has reached a level of performance acceptable to regulators, and genetic tests for warfarin response have not been widely adopted. FitzGerald added that while it is well established that genetic variants impact warfarin metabolism, there is little change in prescribing practices for testing largely because physicians are reluctant to move away from established measures of anticlotting effects. It also still remains to be seen whether there is an impact on clinical outcomes from utilizing this genetic information. Similarly, FitzGerald said, meta-analyses of multiple studies have not suggested a benefit from segregating patients based on genotype in using clopidogrel, which is a medication used to prevent thrombotic events that has a total of $6-$7 billion in annual sales.
Davies stressed the need to consider what the impact on quality and cost of health care will be from genetic or genomic strategies rather than the activity itself. Translating the multitude of genetic and genomic data into a better understanding of disease, improved targets, rapid translation to the clinic, better patient selection, and increased safety will be crucial, he said. FitzGerald noted that “genomic variation is only one hand clapping” and urged that other variables, such as environmental effects, be integrated with genomic information to fully explore the consequences of drug exposure.3 He also suggested that collaboration with sponsors in small studies that utilize next-generation sequencing and drug-evoked phenotyping of adverse events presents an opportunity for genomic and genetic based strategies.
More generally, Ginsburg concluded, the great expectations generated by the promise of genetic-based drug discovery and development have not always been met (Pollack, 2010). According to a recent article in Clinical Pharmacology and Therapeutics, “the level of trust between the different actors in drug development needs to be urgently restored following the disillusionment felt by many that the sequencing of the human genome did
![]()
3 A concept for an integrated data network of genomic and other information is described in Toward Precision Medicine: Building a Knowledge Network for Biomedical Research and a New Taxonomy of Disease (NRC, 2011).
not deliver the expected therapeutic breakthroughs” (Goldman, 2012). But there is optimism moving forward as well, Ginsburg said, noting that the same article pointed out that “there is now a unique window of opportunity to tackle these challenges” and a key model for doing so is establishing new models of collaboration among industry, academia, patient groups, regulators, and biotech companies.












